The evolution of multiple active site configurations in a designed enzyme
- PMID: 30254369
- PMCID: PMC6156567
- DOI: 10.1038/s41467-018-06305-y
The evolution of multiple active site configurations in a designed enzyme
Abstract
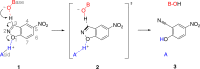
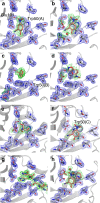
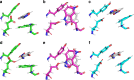
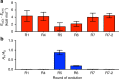
Developments in computational chemistry, bioinformatics, and laboratory evolution have facilitated the de novo design and catalytic optimization of enzymes. Besides creating useful catalysts, the generation and iterative improvement of designed enzymes can provide valuable insight into the interplay between the many phenomena that have been suggested to contribute to catalysis. In this work, we follow changes in conformational sampling, electrostatic preorganization, and quantum tunneling along the evolutionary trajectory of a designed Kemp eliminase. We observe that in the Kemp Eliminase KE07, instability of the designed active site leads to the emergence of two additional active site configurations. Evolutionary conformational selection then gradually stabilizes the most efficient configuration, leading to an improved enzyme. This work exemplifies the link between conformational plasticity and evolvability and demonstrates that residues remote from the active sites of enzymes play crucial roles in controlling and shaping the active site for efficient catalysis.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Evolutionary optimization of computationally designed enzymes: Kemp eliminases of the KE07 series.J Mol Biol. 2010 Mar 5;396(4):1025-42. doi: 10.1016/j.jmb.2009.12.031. Epub 2009 Dec 28. J Mol Biol. 2010. PMID: 20036254
-
Ensemble-based enzyme design can recapitulate the effects of laboratory directed evolution in silico.Nat Commun. 2020 Sep 23;11(1):4808. doi: 10.1038/s41467-020-18619-x. Nat Commun. 2020. PMID: 32968058 Free PMC article.
-
Bridging the gaps in design methodologies by evolutionary optimization of the stability and proficiency of designed Kemp eliminase KE59.Proc Natl Acad Sci U S A. 2012 Jun 26;109(26):10358-63. doi: 10.1073/pnas.1121063109. Epub 2012 Jun 8. Proc Natl Acad Sci U S A. 2012. PMID: 22685214 Free PMC article.
-
Optimization of reorganization energy drives evolution of the designed Kemp eliminase KE07.Biochim Biophys Acta. 2013 May;1834(5):908-17. doi: 10.1016/j.bbapap.2013.01.005. Epub 2013 Feb 1. Biochim Biophys Acta. 2013. PMID: 23380188 Review.
-
The role of reorganization energy in rational enzyme design.Curr Opin Chem Biol. 2014 Aug;21:34-41. doi: 10.1016/j.cbpa.2014.03.011. Epub 2014 Apr 24. Curr Opin Chem Biol. 2014. PMID: 24769299 Review.
Cited by
-
Mutexa: A Computational Ecosystem for Intelligent Protein Engineering.J Chem Theory Comput. 2023 Nov 14;19(21):7459-7477. doi: 10.1021/acs.jctc.3c00602. Epub 2023 Oct 12. J Chem Theory Comput. 2023. PMID: 37828731 Free PMC article. Review.
-
Perspectives on Computational Enzyme Modeling: From Mechanisms to Design and Drug Development.ACS Omega. 2024 Feb 8;9(7):7393-7412. doi: 10.1021/acsomega.3c09084. eCollection 2024 Feb 20. ACS Omega. 2024. PMID: 38405524 Free PMC article. Review.
-
Enzymatic Conjugation of Modified RNA Fragments by Ancestral RNA Ligase AncT4_2.Appl Environ Microbiol. 2022 Dec 13;88(23):e0167922. doi: 10.1128/aem.01679-22. Epub 2022 Nov 23. Appl Environ Microbiol. 2022. PMID: 36416557 Free PMC article.
-
Enhancing a de novo enzyme activity by computationally-focused ultra-low-throughput screening.Chem Sci. 2020 May 19;11(24):6134-6148. doi: 10.1039/d0sc01935f. eCollection 2020 Jun 28. Chem Sci. 2020. PMID: 32832059 Free PMC article.
-
Complete computational design of high-efficiency Kemp elimination enzymes.Nature. 2025 Jul;643(8074):1421-1427. doi: 10.1038/s41586-025-09136-2. Epub 2025 Jun 18. Nature. 2025. PMID: 40533551 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

