Efficient Arrangement of the Replication Fork Trap for In Vitro Propagation of Monomeric Circular DNA in the Chromosome-Replication Cycle Reaction
- PMID: 30257439
- PMCID: PMC6315707
- DOI: 10.3390/life8040043
Efficient Arrangement of the Replication Fork Trap for In Vitro Propagation of Monomeric Circular DNA in the Chromosome-Replication Cycle Reaction
Abstract
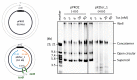
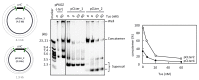
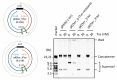
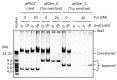
Propagation of genetic information is a fundamental prerequisite for living cells. We recently developed the replication cycle reaction (RCR), an in vitro reaction for circular DNA propagation, by reconstitution of the replication cycle of the Escherichia coli chromosome. In RCR, two replication forks proceed bidirectionally from the replication origin, oriC, and meet at a region opposite oriC, yielding two copies of circular DNA. Although RCR essentially propagates supercoiled monomers, concatemer byproducts are also produced due to inefficient termination of the replication fork progression. Here, we examined the effect of the Tus-ter replication fork trap in RCR. Unexpectedly, when the fork traps were placed opposite oriC, mimicking their arrangement on the chromosome, the propagation of circular DNA was inhibited. On the other hand, fork traps flanking oriC allowed efficient propagation of circular DNA and repressed concatemer production. These findings suggest that collision of the two convergence forks through the fork trap is detrimental to repetition of the replication cycle. We further demonstrate that this detrimental effect was rescued by the UvrD helicase. These results provide insights into the way in which circular DNA monomers replicate repetitively without generating concatemers.
Keywords: DNA amplification; chromosome replication; replication fork; replication termination.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Exponential propagation of large circular DNA by reconstitution of a chromosome-replication cycle.Nucleic Acids Res. 2017 Nov 16;45(20):11525-11534. doi: 10.1093/nar/gkx822. Nucleic Acids Res. 2017. PMID: 29036468 Free PMC article.
-
Chromosomal over-replication in Escherichia coli recG cells is triggered by replication fork fusion and amplified if replichore symmetry is disturbed.Nucleic Acids Res. 2018 Sep 6;46(15):7701-7715. doi: 10.1093/nar/gky566. Nucleic Acids Res. 2018. PMID: 29982635 Free PMC article.
-
An additional replication origin causes cell cycle specific DNA replication fork speed.Front Microbiol. 2025 Apr 30;16:1584664. doi: 10.3389/fmicb.2025.1584664. eCollection 2025. Front Microbiol. 2025. PMID: 40371120 Free PMC article.
-
The replication fork trap and termination of chromosome replication.Mol Microbiol. 2008 Dec;70(6):1323-33. doi: 10.1111/j.1365-2958.2008.06500.x. Epub 2008 Oct 17. Mol Microbiol. 2008. PMID: 19019156 Review.
-
Interplay between chromosomal architecture and termination of DNA replication in bacteria.Front Microbiol. 2023 Jun 26;14:1180848. doi: 10.3389/fmicb.2023.1180848. eCollection 2023. Front Microbiol. 2023. PMID: 37434703 Free PMC article. Review.
Cited by
-
Adaptive evolution of oriC through in vitro propagation of a mini-chromosome in RCR.Nucleic Acids Res. 2025 Aug 11;53(15):gkaf772. doi: 10.1093/nar/gkaf772. Nucleic Acids Res. 2025. PMID: 40795963 Free PMC article.
-
Real-time analysis of initiation regulation systems during the progression of the reconstituted chromosomal replication cycle.Sci Rep. 2025 Feb 8;15(1):4727. doi: 10.1038/s41598-025-88988-0. Sci Rep. 2025. PMID: 39922869 Free PMC article.
-
Building genomes to understand biology.Nat Commun. 2020 Dec 2;11(1):6177. doi: 10.1038/s41467-020-19753-2. Nat Commun. 2020. PMID: 33268788 Free PMC article. Review.
-
A CRISPR-based chromosomal-separation technique for Escherichia coli.Microb Cell Fact. 2022 Nov 11;21(1):235. doi: 10.1186/s12934-022-01957-4. Microb Cell Fact. 2022. PMID: 36369085 Free PMC article.
-
A Fork Trap in the Chromosomal Termination Area Is Highly Conserved across All Escherichia coli Phylogenetic Groups.Int J Mol Sci. 2021 Jul 25;22(15):7928. doi: 10.3390/ijms22157928. Int J Mol Sci. 2021. PMID: 34360694 Free PMC article.
References
-
- Kornberg A., Baker T.A. DNA Replication. 2nd ed. W.H. Freeman; New York, NY, USA: 1992.
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

