Structural basis for broad neutralization of ebolaviruses by an antibody targeting the glycoprotein fusion loop
- PMID: 30258051
- PMCID: PMC6158212
- DOI: 10.1038/s41467-018-06113-4
Structural basis for broad neutralization of ebolaviruses by an antibody targeting the glycoprotein fusion loop
Abstract
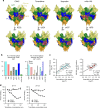
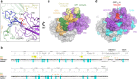
The severity of the 2014-2016 ebolavirus outbreak in West Africa expedited clinical development of therapeutics and vaccines though the countermeasures on hand were largely monospecific and lacked efficacy against other ebolavirus species that previously emerged. Recent studies indicate that ebolavirus glycoprotein (GP) fusion loops are targets for cross-protective antibodies. Here we report the 3.72 Å resolution crystal structure of one such cross-protective antibody, CA45, bound to the ectodomain of Ebola virus (EBOV) GP. The CA45 epitope spans multiple faces of the fusion loop stem, across both GP1 and GP2 subunits, with ~68% of residues identical across > 99.5% of known ebolavirus isolates. Extensive antibody interactions within a pan-ebolavirus small-molecule inhibitor binding cavity on GP define this cavity as a novel site of immune vulnerability. The structure elucidates broad ebolavirus neutralization through a highly conserved epitope on GP and further enables rational design and development of broadly protective vaccines and therapeutics.
Conflict of interest statement
M.J.A. has stocks in Integrated Biotherapeutics, a company developing antibody therapeutics for ebolavirus disease. The remaining authors declare no competing interests.
Figures




Similar articles
-
Structural Basis of Pan-Ebolavirus Neutralization by a Human Antibody against a Conserved, yet Cryptic Epitope.mBio. 2018 Sep 11;9(5):e01674-18. doi: 10.1128/mBio.01674-18. mBio. 2018. PMID: 30206174 Free PMC article.
-
Antibody-Mediated Protective Mechanisms Induced by a Trivalent Parainfluenza Virus-Vectored Ebolavirus Vaccine.J Virol. 2019 Feb 5;93(4):e01845-18. doi: 10.1128/JVI.01845-18. Print 2019 Feb 15. J Virol. 2019. PMID: 30518655 Free PMC article.
-
Mapping of Ebolavirus Neutralization by Monoclonal Antibodies in the ZMapp Cocktail Using Cryo-Electron Tomography and Studies of Cellular Entry.J Virol. 2016 Aug 12;90(17):7618-27. doi: 10.1128/JVI.00406-16. Print 2016 Sep 1. J Virol. 2016. PMID: 27279622 Free PMC article.
-
Structural basis for differential neutralization of ebolaviruses.Viruses. 2012 Apr;4(4):447-70. doi: 10.3390/v4040447. Epub 2012 Apr 5. Viruses. 2012. PMID: 22590681 Free PMC article. Review.
-
Neutralizing ebolavirus: structural insights into the envelope glycoprotein and antibodies targeted against it.Curr Opin Struct Biol. 2009 Aug;19(4):408-17. doi: 10.1016/j.sbi.2009.05.004. Epub 2009 Jun 24. Curr Opin Struct Biol. 2009. PMID: 19559599 Free PMC article. Review.
Cited by
-
Development and Structural Analysis of Antibody Therapeutics for Filoviruses.Pathogens. 2022 Mar 18;11(3):374. doi: 10.3390/pathogens11030374. Pathogens. 2022. PMID: 35335698 Free PMC article. Review.
-
Regions of hepatitis C virus E2 required for membrane association.Nat Commun. 2023 Jan 26;14(1):433. doi: 10.1038/s41467-023-36183-y. Nat Commun. 2023. PMID: 36702826 Free PMC article.
-
The evolution and determinants of neutralization of potent head-binding antibodies against Ebola virus.Cell Rep. 2023 Nov 28;42(11):113366. doi: 10.1016/j.celrep.2023.113366. Epub 2023 Nov 7. Cell Rep. 2023. PMID: 37938974 Free PMC article.
-
Structural Biology Illuminates Molecular Determinants of Broad Ebolavirus Neutralization by Human Antibodies for Pan-Ebolavirus Therapeutic Development.Front Immunol. 2022 Jan 10;12:808047. doi: 10.3389/fimmu.2021.808047. eCollection 2021. Front Immunol. 2022. PMID: 35082794 Free PMC article. Review.
-
Designs and Characterization of Subunit Ebola GP Vaccine Candidates: Implications for Immunogenicity.Front Immunol. 2020 Nov 4;11:586595. doi: 10.3389/fimmu.2020.586595. eCollection 2020. Front Immunol. 2020. PMID: 33250896 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

