Nuclear lamina dysfunction triggers a germline stem cell checkpoint
- PMID: 30262885
- PMCID: PMC6160405
- DOI: 10.1038/s41467-018-06277-z
Nuclear lamina dysfunction triggers a germline stem cell checkpoint
Abstract
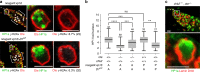
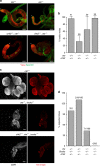
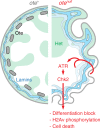
LEM domain (LEM-D) proteins are conserved components of the nuclear lamina (NL) that contribute to stem cell maintenance through poorly understood mechanisms. The Drosophila emerin homolog Otefin (Ote) is required for maintenance of germline stem cells (GSCs) and gametogenesis. Here, we show that ote mutants carry germ cell-specific changes in nuclear architecture that are linked to GSC loss. Strikingly, we found that both GSC death and gametogenesis are rescued by inactivation of the DNA damage response (DDR) kinases, ATR and Chk2. Whereas the germline checkpoint draws from components of the DDR pathway, genetic and cytological features of the GSC checkpoint differ from the canonical pathway. Instead, structural deformation of the NL correlates with checkpoint activation. Despite remarkably normal oogenesis, rescued oocytes do not support embryogenesis. Taken together, these data suggest that NL dysfunction caused by Otefin loss triggers a GSC-specific checkpoint that contributes to maintenance of gamete quality.
Conflict of interest statement
The authors declare no competing interests.
Figures

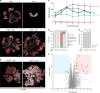
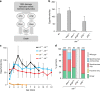
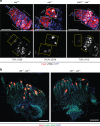
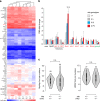
Similar articles
-
Survival of Drosophila germline stem cells requires the chromatin-binding protein Barrier-to-autointegration factor.Development. 2020 May 13;147(9):dev186171. doi: 10.1242/dev.186171. Development. 2020. PMID: 32345742 Free PMC article.
-
Drosophila male and female germline stem cell niches require the nuclear lamina protein Otefin.Dev Biol. 2016 Jul 1;415(1):75-86. doi: 10.1016/j.ydbio.2016.05.001. Epub 2016 May 10. Dev Biol. 2016. PMID: 27174470 Free PMC article.
-
Emerin preserves stem cell survival through maintenance of centrosome and nuclear lamina structure.Development. 2024 Nov 15;151(22):dev204219. doi: 10.1242/dev.204219. Epub 2024 Nov 13. Development. 2024. PMID: 39465887 Free PMC article.
-
Nuclear architecture as an intrinsic regulator of Drosophila female germline stem cell maintenance.Curr Opin Insect Sci. 2020 Feb;37:30-38. doi: 10.1016/j.cois.2019.11.007. Epub 2020 Jan 30. Curr Opin Insect Sci. 2020. PMID: 32087561 Free PMC article. Review.
-
Germline stem cells.Curr Top Dev Biol. 2013;102:97-126. doi: 10.1016/B978-0-12-416024-8.00004-0. Curr Top Dev Biol. 2013. PMID: 23287031 Review.
Cited by
-
Nuclear Envelope Membrane Protein 1 plays crucial and conserved roles in female meiosis.Res Sq [Preprint]. 2025 Aug 4:rs.3.rs-7159889. doi: 10.21203/rs.3.rs-7159889/v1. Res Sq. 2025. PMID: 40799732 Free PMC article. Preprint.
-
Germ Cell Lineage Homeostasis in Drosophila Requires the Vasa RNA Helicase.Genetics. 2019 Nov;213(3):911-922. doi: 10.1534/genetics.119.302558. Epub 2019 Sep 4. Genetics. 2019. PMID: 31484689 Free PMC article.
-
Survival of Drosophila germline stem cells requires the chromatin-binding protein Barrier-to-autointegration factor.Development. 2020 May 13;147(9):dev186171. doi: 10.1242/dev.186171. Development. 2020. PMID: 32345742 Free PMC article.
-
Developmental changes in nuclear lamina components during germ cell differentiation.Nucleus. 2024 Dec;15(1):2339214. doi: 10.1080/19491034.2024.2339214. Epub 2024 Apr 10. Nucleus. 2024. PMID: 38597409 Free PMC article.
-
Maintenance of germline stem cell homeostasis despite severe nuclear distortion.Dev Biol. 2024 Nov;515:139-150. doi: 10.1016/j.ydbio.2024.07.009. Epub 2024 Jul 20. Dev Biol. 2024. PMID: 39038593
References
Publication types
MeSH terms
Substances
Grants and funding
- R01 GM087341/GM/NIGMS NIH HHS/United States
- S10 RR025439/RR/NCRR NIH HHS/United States
- P30 CA086862/CA/NCI NIH HHS/United States
- GM087341/U.S. Department of Health & Human Services | NIH | National Institute of General Medical Sciences (NIGMS)/International
- DRG-2185-14/Damon Runyon Cancer Research Foundation (Damon Runyon)/International
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

