The Origin of Mitochondria-Specific Outer Membrane β-Barrels from an Ancestral Bacterial Fragment
- PMID: 30265295
- PMCID: PMC6193526
- DOI: 10.1093/gbe/evy216
The Origin of Mitochondria-Specific Outer Membrane β-Barrels from an Ancestral Bacterial Fragment
Abstract
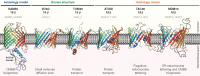
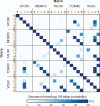
Outer membrane β-barrels (OMBBs) are toroidal arrays of antiparallel β-strands that span the outer membrane of Gram-negative bacteria and eukaryotic organelles. Although homologous, most families of bacterial OMBBs evolved through the independent amplification of an ancestral ββ-hairpin. In mitochondria, one family (SAM50) has a clear bacterial ancestry; the origin of the other family, consisting of 19-stranded OMBBs found only in mitochondria (MOMBBs), is substantially unclear. In a large-scale comparison of mitochondrial and bacterial OMBBs, we find evidence that the common ancestor of all MOMBBs emerged by the amplification of a double ββ-hairpin of bacterial origin, probably at the time of the Last Eukaryotic Common Ancestor. Thus, MOMBBs are indeed descended from bacterial OMBBs, but their fold formed independently in the proto-mitochondria, possibly in response to the need for a general-purpose polypeptide importer. This occurred by a process of amplification, despite the final fold having a prime number of strands.
Figures



Similar articles
-
Evolution of outer membrane beta-barrels from an ancestral beta beta hairpin.Mol Biol Evol. 2010 Jun;27(6):1348-58. doi: 10.1093/molbev/msq017. Epub 2010 Jan 27. Mol Biol Evol. 2010. PMID: 20106904
-
Gram-negative outer-membrane proteins with multiple β-barrel domains.Proc Natl Acad Sci U S A. 2021 Aug 3;118(31):e2104059118. doi: 10.1073/pnas.2104059118. Proc Natl Acad Sci U S A. 2021. PMID: 34330833 Free PMC article.
-
Structure and evolution of mitochondrial outer membrane proteins of beta-barrel topology.Biochim Biophys Acta. 2010 Jun-Jul;1797(6-7):1292-9. doi: 10.1016/j.bbabio.2010.04.019. Epub 2010 May 5. Biochim Biophys Acta. 2010. PMID: 20450883 Review.
-
Biogenesis of beta-barrel membrane proteins of mitochondria.Trends Biochem Sci. 2005 Oct;30(10):575-82. doi: 10.1016/j.tibs.2005.08.009. Trends Biochem Sci. 2005. PMID: 16126389 Review.
-
Outer membrane β-barrel structure prediction through the lens of AlphaFold2.Proteins. 2024 Jan;92(1):3-14. doi: 10.1002/prot.26552. Epub 2023 Jul 19. Proteins. 2024. PMID: 37465978
Cited by
-
Structural diversity of oligomeric β-propellers with different numbers of identical blades.Elife. 2019 Oct 15;8:e49853. doi: 10.7554/eLife.49853. Elife. 2019. PMID: 31613220 Free PMC article.
-
The Evolutionary History of Peptidases Involved in the Processing of Organelle-Targeting Peptides.Genome Biol Evol. 2022 Jul 2;14(7):evac101. doi: 10.1093/gbe/evac101. Genome Biol Evol. 2022. PMID: 35758251 Free PMC article.
-
Historical Perspective of Pore-Forming Activity Studies of Voltage-Dependent Anion Channel (Eukaryotic or Mitochondrial Porin) Since Its Discovery in the 70th of the Last Century.Front Physiol. 2021 Oct 26;12:734226. doi: 10.3389/fphys.2021.734226. eCollection 2021. Front Physiol. 2021. PMID: 35547863 Free PMC article. Review.
-
The Name Is Barrel, β-Barrel.Methods Mol Biol. 2024;2778:1-30. doi: 10.1007/978-1-0716-3734-0_1. Methods Mol Biol. 2024. PMID: 38478268
-
Sam50 exerts neuroprotection by maintaining the mitochondrial structure during experimental cerebral ischemia/reperfusion injury in rats.CNS Neurosci Ther. 2022 Dec;28(12):2230-2244. doi: 10.1111/cns.13967. Epub 2022 Sep 8. CNS Neurosci Ther. 2022. PMID: 36074556 Free PMC article.
References
-
- Alva V, Lupas AN.. 2018. From ancestral peptides to designed proteins. Curr Opin Struct Biol. 48:103–109. - PubMed
-
- Andersson SGE, et al. 1998. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396(6707):133–140. - PubMed
-
- Andrade MA, Perez-Iratxeta C, Ponting CP.. 2001. Protein repeats: structures, functions, and evolution. J Struct Biol. 134(2–3):117–131. - PubMed
-
- Bay DC, Hafez M, Young MJ, Court DA.. 2012. Phylogenetic and coevolutionary analysis of the β-barrel protein family comprised of mitochondrial porin (VDAC) and Tom40. Biochim Biophys Acta – Biomembr. 1818(6):1502–1519. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

