Machine Learning Approaches for Protein⁻Protein Interaction Hot Spot Prediction: Progress and Comparative Assessment
- PMID: 30287797
- PMCID: PMC6222875
- DOI: 10.3390/molecules23102535
Machine Learning Approaches for Protein⁻Protein Interaction Hot Spot Prediction: Progress and Comparative Assessment
Abstract
Hot spots are the subset of interface residues that account for most of the binding free energy, and they play essential roles in the stability of protein binding. Effectively identifying which specific interface residues of protein⁻protein complexes form the hot spots is critical for understanding the principles of protein interactions, and it has broad application prospects in protein design and drug development. Experimental methods like alanine scanning mutagenesis are labor-intensive and time-consuming. At present, the experimentally measured hot spots are very limited. Hence, the use of computational approaches to predicting hot spots is becoming increasingly important. Here, we describe the basic concepts and recent advances of machine learning applications in inferring the protein⁻protein interaction hot spots, and assess the performance of widely used features, machine learning algorithms, and existing state-of-the-art approaches. We also discuss the challenges and future directions in the prediction of hot spots.
Keywords: hot spots; machine learning; performance evaluation; protein-protein interaction.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
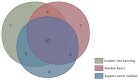
References
-
- Zeng J., Li D., Wu Y., Zou Q., Liu X. An empirical study of features fusion techniques for protein-protein interaction prediction. Curr. Bioinform. 2016;11:4–12. doi: 10.2174/1574893611666151119221435. - DOI
-
- Fischer T., Arunachalam K., Bailey D., Mangual V., Bakhru S., Russo R., Huang D., Paczkowski M., Lalchandani V., Ramachandra C., et al. The binding interface database (BID): a compilation of amino acid hot spots in protein interfaces. Bioinformatics. 2003;19:1453–1454. doi: 10.1093/bioinformatics/btg163. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

