Engineered dCas9 with reduced toxicity in bacteria: implications for genetic circuit design
- PMID: 30289463
- PMCID: PMC6237744
- DOI: 10.1093/nar/gky884
Engineered dCas9 with reduced toxicity in bacteria: implications for genetic circuit design
Abstract
Large synthetic genetic circuits require the simultaneous expression of many regulators. Deactivated Cas9 (dCas9) can serve as a repressor by having a small guide RNA (sgRNA) direct it to bind a promoter. The programmability and specificity of RNA:DNA basepairing simplifies the generation of many orthogonal sgRNAs that, in theory, could serve as a large set of regulators in a circuit. However, dCas9 is toxic in many bacteria, thus limiting how high it can be expressed, and low concentrations are quickly sequestered by multiple sgRNAs. Here, we construct a non-toxic version of dCas9 by eliminating PAM (protospacer adjacent motif) binding with a R1335K mutation (dCas9*) and recovering DNA binding by fusing it to the PhlF repressor (dCas9*_PhlF). Both the 30 bp PhlF operator and 20 bp sgRNA binding site are required to repress a promoter. The larger region required for recognition mitigates toxicity in Escherichia coli, allowing up to 9600 ± 800 molecules of dCas9*_PhlF per cell before growth or morphology are impacted, as compared to 530 ± 40 molecules of dCas9. Further, PhlF multimerization leads to an increase in average cooperativity from n = 0.9 (dCas9) to 1.6 (dCas9*_PhlF). A set of 30 orthogonal sgRNA-promoter pairs are characterized as NOT gates; however, the simultaneous use of multiple sgRNAs leads to a monotonic decline in repression and after 15 are co-expressed the dynamic range is <10-fold. This work introduces a non-toxic variant of dCas9, critical for its use in applications in metabolic engineering and synthetic biology, and exposes a limitation in the number of regulators that can be used in one cell when they rely on a shared resource.
Figures


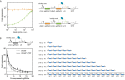
References
-
- Purnick P.E., Weiss R.. The second wave of synthetic biology: from modules to systems. Nat. Rev. Mol. Cell. Biol. 2009; 10:410. - PubMed
-
- Nielsen A.A.K., Segall-Shapiro T.H., Voigt C.A.. Advances in genetic circuit design: novel biochemistries, deep part mining, and precision gene expression. Curr. Opin. Chem. Biol. 2013; 17:878–892. - PubMed
-
- Gaber R., Lebar T., Majerle A., Šter B., Dobnikar A., Benčina M., Jerala R.. Designable DNA-binding domains enable construction of logic circuits in mammalian cells. Nat. Chem. Biol. 2014; 10:203. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

