Structural basis for targeting avian sarcoma virus Gag polyprotein to the plasma membrane for virus assembly
- PMID: 30309983
- PMCID: PMC6295720
- DOI: 10.1074/jbc.RA118.003944
Structural basis for targeting avian sarcoma virus Gag polyprotein to the plasma membrane for virus assembly
Abstract
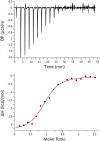
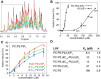
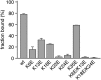
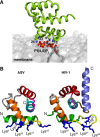
For most retroviruses, including HIV-1, binding of the Gag polyprotein to the plasma membrane (PM) is mediated by interactions between Gag's N-terminal myristoylated matrix (MA) domain and phosphatidylinositol 4,5-bisphosphate (PI(4,5)P2) in the PM. The Gag protein of avian sarcoma virus (ASV) lacks the N-myristoylation signal but contains structural domains having functions similar to those of HIV-1 Gag. The molecular mechanism by which ASV Gag binds to the PM is incompletely understood. Here, we employed NMR techniques to elucidate the molecular determinants of the membrane-binding domain of ASV MA (MA87) to lipids and liposomes. We report that MA87 binds to the polar head of phosphoinositides such as PI(4,5)P2 We found that MA87 binding to inositol phosphates (IPs) is significantly enhanced by increasing the number of phosphate groups, indicating that the MA87-IP binding is governed by charge-charge interactions. Using a sensitive NMR-based liposome-binding assay, we show that binding of MA87 to liposomes is enhanced by incorporation of PI(4,5)P2 and phosphatidylserine. We also show that membrane binding is mediated by a basic surface formed by Lys-6, Lys-13, Lys-23, and Lys-24. Substitution of these residues to glutamate abolished binding of MA87 to both IPs and liposomes. In an accompanying paper, we further report that mutation of these lysine residues diminishes Gag assembly on the PM and inhibits ASV particle release. These findings provide a molecular basis for ASV Gag binding to the inner leaflet of the PM and advance our understanding of the basic mechanisms of retroviral assembly.
Keywords: ASV; Gag protein; avian sarcoma virus; human immunodeficiency virus (HIV); inositol hexakisphosphate; isothermal titration calorimetry (ITC); liposome; myristoylated matrix; phosphatidylinositol 4,5-bisphosphate; phosphatidylserine; plasma membrane.
© 2018 Vlach et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures




References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

