Enzymatic complexes across scales
- PMID: 30315098
- PMCID: PMC6204551
- DOI: 10.1042/EBC20180008
Enzymatic complexes across scales
Abstract
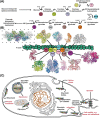
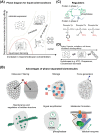
An unprecedented opportunity to integrate ~100 years of meticulous in vitro biomolecular research is currently provided in the light of recent advances in methods to visualize closer-to-native architectures of biomolecular machines, and metabolic enzymes in particular. Traditional views of enzymes, namely biomolecular machines, only partially explain their role, organization and kinetics in the cellular milieu. Enzymes self- or hetero-associate, form fibers, may bind to membranes or cytoskeletal elements, have regulatory roles, associate into higher order assemblies (metabolons) or even actively participate in phase-separated membraneless organelles, and all the above in a transient, temporal and spatial manner in response to environmental changes or structural/functional changes of their assemblies. Here, we focus on traditional and emerging concepts in cellular biochemistry and discuss new opportunities in bridging structural, molecular and cellular analyses for metabolic pathways, accumulated over the years, highlighting functional aspects of enzymatic complexes discussed across different levels of spatial resolution.
Keywords: electron microscopy; enzymology; glycolysis; protein-protein interactions.
© 2018 The Author(s).
Conflict of interest statement
The authors declare that there are no competing interests associated with the manuscript.
Figures

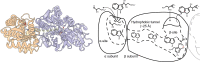

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

