The Long Non-coding RNA Flatr Anticipates Foxp3 Expression in Regulatory T Cells
- PMID: 30319599
- PMCID: PMC6167443
- DOI: 10.3389/fimmu.2018.01989
The Long Non-coding RNA Flatr Anticipates Foxp3 Expression in Regulatory T Cells
Abstract
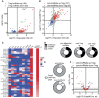
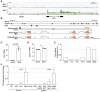
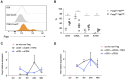
Mammalian genomes encode a plethora of long non-coding RNA (lncRNA). These transcripts are thought to regulate gene expression, influencing biological processes from development to pathology. Results from the few lncRNA that have been studied in the context of the immune system have highlighted potentially critical functions as network regulators. Here we explored the nature of the lncRNA transcriptome in regulatory T cells (Tregs), a subset of CD4+ T cells required to establish and maintain immunological self-tolerance. The identified Treg lncRNA transcriptome showed distinct differences from that of non-regulatory CD4+ T cells, with evidence of direct shaping of the lncRNA transcriptome by Foxp3, the master transcription factor driving the distinct mRNA profile of Tregs. Treg lncRNA changes were disproportionally reversed in the absence of Foxp3, with an enrichment for colocalisation with Foxp3 DNA binding sites, indicating a direct coordination of transcription by Foxp3 independent of the mRNA coordination function. We further identified a novel lncRNA Flatr, as a member of the core Treg lncRNA transcriptome. Flatr expression anticipates Foxp3 expression during in vitro Treg conversion, and Flatr-deficient mice show a mild delay in in vitro and peripheral Treg induction. These results implicate Flatr as part of the upstream cascade leading to Treg conversion, and may provide clues as to the nature of this process.
Keywords: Flatr; Foxp3; Tregs; iTreg induction; lncRNA.
Figures




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

