Microarray analysis reveals long non‑coding RNA SOX2OT as a novel candidate regulator in diabetic nephropathy
- PMID: 30320339
- PMCID: PMC6236268
- DOI: 10.3892/mmr.2018.9534
Microarray analysis reveals long non‑coding RNA SOX2OT as a novel candidate regulator in diabetic nephropathy
Abstract
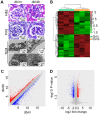
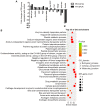
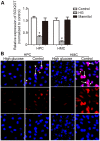
Diabetic nephropathy (DN) is a highly complex syndrome involving multiple dysregulated biological processes. Long non‑coding RNAs (lncRNAs) are now believed to have an important function in various diseases. However, their roles in DN remain largely unknown. Therefore, the present study was performed in order to investigate the lncRNAs that have a crucial role in DN. db/db mice were used as a DN model while db/m mice served as a control to search for lncRNAs which may have important roles in DN. Microarray and bioinformatics analysis gave an overview of the features of differentially expressed genes. Gene Ontology and Kyoto Encyclopedia of Genes and Genomes enrichment analysis demonstrated the typical biological alterations in DN. A co‑expression network of lncRNAs and mRNAs revealed the complex interaction pattern in DN conditions. Further data investigation indicated that SOX2‑overlapping transcript (SOX2OT), which was significantly downregulated in DN mice, may be the potentially functional lncRNA contributing to the onset of DN. The UCSC database demonstrated that SOX2OT was highly conserved in mice and humans. Additionally further study using cultured human podocytes and mesangial cells confirmed the downregulation of SOX2OT using reverse transcription‑quantitative polymerase chain reaction and fluorescence in situ hybridization. However, the cellular location of SOX2OT depended on certain cell types. Taken together, the results of the present study indicated that SOX2OT may act as an important regulator in the pathogenesis of DN by interacting with various mRNAs with critical roles in DN.
Keywords: SOX2-overlapping transcript; bioinformatics analysis; microarray analysis; diabetic nephropathy.
Figures



References
-
- Shang J, Wan Q, Wang X, Duan Y, Wang Z, Wei X, Zhang Y, Wang H, Wang R, Yi F. Identification of NOD2 as a novel target of RNA-binding protein HuR: Evidence from NADPH oxidase-mediated HuR signaling in diabetic nephropathy. Free Radic Biol Med. 2015;79:217–227. doi: 10.1016/j.freeradbiomed.2014.12.013. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

