Alcohol Dehydrogenases, Aldehyde Dehydrogenases, and Alcohol Use Disorders: A Critical Review
- PMID: 30320893
- PMCID: PMC6286250
- DOI: 10.1111/acer.13904
Alcohol Dehydrogenases, Aldehyde Dehydrogenases, and Alcohol Use Disorders: A Critical Review
Abstract
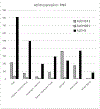
Alcohol use disorders (AUDs) are complex traits, meaning that variations in many genes contribute to the risk, as does the environment. Although the total genetic contribution to risk is substantial, most individual variations make only very small contributions. By far the strongest contributors are functional variations in 2 genes involved in alcohol (ethanol [EtOH]) metabolism. A functional variant in alcohol dehydrogenase 1B (ADH1B) is protective in people of European and Asian descent, and a different functional variant in the same gene is protective in those of African descent. A strongly protective variant in aldehyde dehydrogenase 2 (ALDH2) is essentially only found in Asians. This highlights the need to study a wide range of populations. The likely mechanism of protection against heavy drinking and AUDs in both cases is alteration in the rate of metabolism of EtOH that at least transiently elevates acetaldehyde. Other ADH and ALDH variants, including functional variations in ADH1C, have also been implicated in affecting drinking behavior and risk for alcoholism. The pattern of linkage disequilibrium in the ADH region and the differences among populations complicate analyses, particularly of regulatory variants. This critical review focuses upon the ADH and ALDH genes as they affect AUDs.
Keywords: ADH Genes; ALDH Genes; Alcohol Consumption; Alcohol Dependence; Alcohol Metabolism; Linkage Disequilibrium.
© 2018 by the Research Society on Alcoholism.
Figures
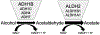

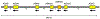
References
-
- Adachi J, Mizoi Y, Fukunaga T, Ogawa Y, Imamichi H (1989) Comparative study on ethanol elimination and blood acetaldehyde between alcoholics and control subjects. Alcohol Clin Exp Res 13:601–604. - PubMed
-
- American Psychiatric Association (2013) Diagnostic and Statistical Manual of Mental Disorders: DSM-5, AMERICAN PSYCHIATRIC PUBLISHING.
-
- Baik I, Cho NH, Kim SH, Han BG, Shin C (2011) Genome-wide association studies identify genetic loci related to alcohol consumption in Korean men. Am J Clin Nutr 93:809–816. - PubMed
-
- Biernacka JM, Geske JR, Schneekloth TD, Frye MA, Cunningham JM, Choi DS, Tapp CL, Lewis BR, Drews MS, T LP, Colby CL, Hall-Flavin DK, Loukianova LL, Heit JA, Mrazek DA, Karpyak VM (2013) Replication of genome wide association studies of alcohol dependence: support for association with variation in ADH1C. PLoS One 8:e58798. - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

