Altered Levels of Long NcRNAs Meg3 and Neat1 in Cell And Animal Models Of Huntington's Disease
- PMID: 30321100
- PMCID: PMC6284602
- DOI: 10.1080/15476286.2018.1534524
Altered Levels of Long NcRNAs Meg3 and Neat1 in Cell And Animal Models Of Huntington's Disease
Abstract
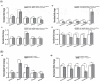
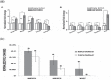
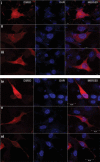
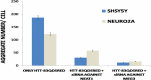
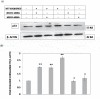
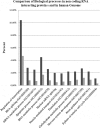
Altered expression levels of protein-coding genes and microRNAs have been implicated in the pathogenesis of Huntington's disease (HD). The involvement of other ncRNAs, especially long ncRNAs (lncRNA), is being realized recently and the related knowledge is still rudimentary. Using small RNA sequencing and PCR arrays we observed perturbations in the levels of 12 ncRNAs in HD mouse brain, eight of which had human homologs. Of these, Meg3, Neat1, and Xist showed a consistent and significant increase in HD cell and animal models. Transient knock-down of Meg3 and Neat1 in cell models of HD led to a significant decrease of aggregates formed by mutant huntingtin and downregulation of the endogenous Tp53 expression. Understanding Meg3 and Neat1 functions in the context of HD pathogenesis is likely to open up new strategies to control the disease.
Keywords: Huntington’s disease; MEG3; NEAT1; long ncRNA.
Figures

Similar articles
-
Imaging and Assay of the Dynamics of Cytotoxic Huntingtin (HTT) Protein Aggregates Regulated by lncRNAs.Methods Mol Biol. 2024;2761:421-430. doi: 10.1007/978-1-0716-3662-6_29. Methods Mol Biol. 2024. PMID: 38427253
-
Dysregulation of long non-coding RNAs and their mechanisms in Huntington's disease.J Neurosci Res. 2021 Sep;99(9):2074-2090. doi: 10.1002/jnr.24825. Epub 2021 May 24. J Neurosci Res. 2021. PMID: 34031910 Review.
-
Altered Expression of the Long Noncoding RNA NEAT1 in Huntington's Disease.Mol Neurobiol. 2017 Mar;54(2):1577-1586. doi: 10.1007/s12035-016-9928-9. Epub 2016 May 25. Mol Neurobiol. 2017. PMID: 27221610
-
Downregulation of glial genes involved in synaptic function mitigates Huntington's disease pathogenesis.Elife. 2021 Apr 19;10:e64564. doi: 10.7554/eLife.64564. Elife. 2021. PMID: 33871358 Free PMC article.
-
Long non-coding RNAs in Huntington's disease neurodegeneration.Neurobiol Dis. 2012 May;46(2):245-54. doi: 10.1016/j.nbd.2011.12.006. Epub 2011 Dec 14. Neurobiol Dis. 2012. PMID: 22202438 Review.
Cited by
-
Integrated analysis of differentially expressed genes and construction of a competing endogenous RNA network in human Huntington neural progenitor cells.BMC Med Genomics. 2021 Feb 12;14(1):48. doi: 10.1186/s12920-021-00894-2. BMC Med Genomics. 2021. PMID: 33579286 Free PMC article.
-
Molecular landscape of long noncoding RNAs in brain disorders.Mol Psychiatry. 2021 Apr;26(4):1060-1074. doi: 10.1038/s41380-020-00947-5. Epub 2020 Nov 10. Mol Psychiatry. 2021. PMID: 33173194 Review.
-
The emerging role of long non-coding RNAs, microRNAs, and an accelerated epigenetic age in Huntington's disease.Front Aging Neurosci. 2022 Sep 15;14:987174. doi: 10.3389/fnagi.2022.987174. eCollection 2022. Front Aging Neurosci. 2022. PMID: 36185471 Free PMC article. Review.
-
Long non-coding RNA RP11-283G6.5 confines breast cancer development through modulating miR-188-3p/TMED3/Wnt/β-catenin signalling.RNA Biol. 2021 Oct 15;18(sup1):287-302. doi: 10.1080/15476286.2021.1941608. Epub 2021 Jul 27. RNA Biol. 2021. PMID: 34130584 Free PMC article.
-
Current Diagnostic Methods and Non-Coding RNAs as Possible Biomarkers in Huntington's Disease.Genes (Basel). 2022 Nov 3;13(11):2017. doi: 10.3390/genes13112017. Genes (Basel). 2022. PMID: 36360254 Free PMC article. Review.
References
-
- Huntington’s Disease Collaborative Research Group A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell. 1993;72:971–983. - PubMed
-
- Imarisio S, Carmichael J, Korolchuk V, et al. Huntington’s disease: from pathology and genetics to potential therapies. Biochem J. 2008;412:191–209. - PubMed
-
- Seredenina T, Luthi-Carter R. What have we learned from gene expression profiles in Huntington’s disease? Neurobiol Dis. 2012;45:83–98. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous
