Proteomic profiling and functional characterization of post-translational modifications of the fission yeast RNA exosome
- PMID: 30321377
- PMCID: PMC6265454
- DOI: 10.1093/nar/gky915
Proteomic profiling and functional characterization of post-translational modifications of the fission yeast RNA exosome
Abstract
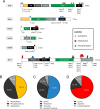
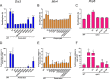
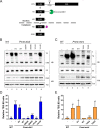
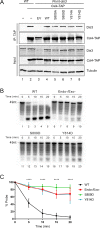
The RNA exosome is a conserved multi-subunit complex essential for processing and degradation of several types of RNAs. Although many of the functions of the RNA exosome are well established, whether the activity of this complex is regulated remains unclear. Here we performed a proteomic analysis of the RNA exosome complex purified from Schizosaccharomyces pombe and identified 39 post-translational modifications (PTMs), including phosphorylation, methylation, and acetylation sites. Interestingly, most of the modifications were identified in Dis3, a catalytic subunit of the RNA exosome, as well as in the exosome-associated RNA helicase, Mtr4. Functional analysis of selected PTM sites using modification-deficient and -mimetic versions of exosome subunits revealed substitutions that affected cell growth and exosome functions. Notably, our results suggest that site-specific phosphorylation in the catalytic center of Dis3 and in the helical bundle domain of Mtr4 control their activity. Our findings support a view in which post-translational modifications fine-tune exosome activity and add a layer of regulation to RNA degradation.
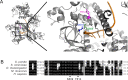
Figures

Similar articles
-
Helicase-Dependent RNA Decay Illuminated by a Cryo-EM Structure of a Human Nuclear RNA Exosome-MTR4 Complex.Cell. 2018 Jun 14;173(7):1663-1677.e21. doi: 10.1016/j.cell.2018.05.041. Cell. 2018. PMID: 29906447 Free PMC article.
-
Structure and reconstitution of yeast Mpp6-nuclear exosome complexes reveals that Mpp6 stimulates RNA decay and recruits the Mtr4 helicase.Elife. 2017 Jul 25;6:e29062. doi: 10.7554/eLife.29062. Elife. 2017. PMID: 28742025 Free PMC article.
-
Reconstitution of the Schizosaccharomyces pombe RNA Exosome.Methods Mol Biol. 2020;2062:449-465. doi: 10.1007/978-1-4939-9822-7_22. Methods Mol Biol. 2020. PMID: 31768990 Free PMC article.
-
The Nuclear RNA Exosome and Its Cofactors.Adv Exp Med Biol. 2019;1203:113-132. doi: 10.1007/978-3-030-31434-7_4. Adv Exp Med Biol. 2019. PMID: 31811632 Review.
-
Comparison of the yeast and human nuclear exosome complexes.Biochem Soc Trans. 2012 Aug;40(4):850-5. doi: 10.1042/BST20120061. Biochem Soc Trans. 2012. PMID: 22817747 Review.
Cited by
-
Non-mitochondrial aconitase regulates the expression of iron-uptake genes by controlling the RNA turnover process in fission yeast.J Microbiol. 2021 Dec;59(12):1075-1082. doi: 10.1007/s12275-021-1438-4. Epub 2021 Oct 26. J Microbiol. 2021. PMID: 34705258
-
RNA-binding protein Mub1 and the nuclear RNA exosome act to fine-tune environmental stress response.Life Sci Alliance. 2021 Nov 30;5(2):e202101111. doi: 10.26508/lsa.202101111. Print 2022 Feb. Life Sci Alliance. 2021. PMID: 34848435 Free PMC article.
-
Mtr4 RNA helicase structures and interactions.Biol Chem. 2021 Jan 6;402(5):605-616. doi: 10.1515/hsz-2020-0329. Print 2021 Apr 27. Biol Chem. 2021. PMID: 33857361 Free PMC article. Review.
-
The biology of exosomes and exosomal non-coding RNAs in cardiovascular diseases.Front Pharmacol. 2025 May 26;16:1529375. doi: 10.3389/fphar.2025.1529375. eCollection 2025. Front Pharmacol. 2025. PMID: 40492132 Free PMC article. Review.
-
Yeast Genome Maintenance by the Multifunctional PIF1 DNA Helicase Family.Genes (Basel). 2020 Feb 20;11(2):224. doi: 10.3390/genes11020224. Genes (Basel). 2020. PMID: 32093266 Free PMC article. Review.
References
-
- Andrulis E.D., Werner J., Nazarian A., Erdjument-Bromage H., Tempst P., Lis J.T.. The RNA processing exosome is linked to elongating RNA polymerase II in Drosophila. Nature. 2002; 420:837–841. - PubMed
-
- Chekanova J.A., Gregory B.D., Reverdatto S.V., Chen H., Kumar R., Hooker T., Yazaki J., Li P., Skiba N., Peng Q. et al. . Genome-wide high-resolution mapping of exosome substrates reveals hidden features in the Arabidopsis transcriptome. Cell. 2007; 131:1340–1353. - PubMed
-
- Lemay J.F., D’Amours A., Lemieux C., Lackner D.H., St-Sauver V.G., Bahler J., Bachand F.. The nuclear poly(A)-binding protein interacts with the exosome to promote synthesis of noncoding small nucleolar RNAs. Mol. Cell. 2010; 37:34–45. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

