Dynamic regulation of HIV-1 capsid interaction with the restriction factor TRIM5α identified by magic-angle spinning NMR and molecular dynamics simulations
- PMID: 30333189
- PMCID: PMC6233135
- DOI: 10.1073/pnas.1800796115
Dynamic regulation of HIV-1 capsid interaction with the restriction factor TRIM5α identified by magic-angle spinning NMR and molecular dynamics simulations
Abstract
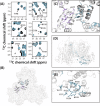
The host factor protein TRIM5α plays an important role in restricting the host range of HIV-1, interfering with the integrity of the HIV-1 capsid. TRIM5 triggers an antiviral innate immune response by functioning as a capsid pattern recognition receptor, although the precise mechanism by which the restriction is imposed is not completely understood. Here we used an integrated magic-angle spinning nuclear magnetic resonance and molecular dynamics simulations approach to characterize, at atomic resolution, the dynamics of the capsid's hexameric and pentameric building blocks, and the interactions with TRIM5α in the assembled capsid. Our data indicate that assemblies in the presence of the pentameric subunits are more rigid on the microsecond to millisecond timescales than tubes containing only hexamers. This feature may be of key importance for controlling the capsid's morphology and stability. In addition, we found that TRIM5α binding to capsid induces global rigidification and perturbs key intermolecular interfaces essential for higher-order capsid assembly, with structural and dynamic changes occurring throughout the entire CA polypeptide chain in the assembly, rather than being limited to a specific protein-protein interface. Taken together, our results suggest that TRIM5α uses several mechanisms to destabilize the capsid lattice, ultimately inducing its disassembly. Our findings add to a growing body of work indicating that dynamic allostery plays a pivotal role in capsid assembly and HIV-1 infectivity.
Keywords: CA protein assemblies; HIV-1 capsid; HIV-AIDS; TRIM5α; magic-angle spinning NMR.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
General Model for Retroviral Capsid Pattern Recognition by TRIM5 Proteins.J Virol. 2018 Jan 30;92(4):e01563-17. doi: 10.1128/JVI.01563-17. Print 2018 Feb 15. J Virol. 2018. PMID: 29187540 Free PMC article.
-
The Three-Fold Axis of the HIV-1 Capsid Lattice Is the Species-Specific Binding Interface for TRIM5α.J Virol. 2018 Feb 12;92(5):e01541-17. doi: 10.1128/JVI.01541-17. Print 2018 Mar 1. J Virol. 2018. PMID: 29237846 Free PMC article.
-
Rhesus monkey TRIM5α SPRY domain recognizes multiple epitopes that span several capsid monomers on the surface of the HIV-1 mature viral core.J Mol Biol. 2013 Dec 13;425(24):5032-44. doi: 10.1016/j.jmb.2013.07.025. Epub 2013 Jul 23. J Mol Biol. 2013. PMID: 23886867 Free PMC article.
-
Molecular Architecture of the Retroviral Capsid.Trends Biochem Sci. 2016 May;41(5):410-420. doi: 10.1016/j.tibs.2016.02.009. Epub 2016 Mar 30. Trends Biochem Sci. 2016. PMID: 27039020 Free PMC article. Review.
-
Visualizing HIV-1 Capsid and Its Interactions with Antivirals and Host Factors.Viruses. 2021 Feb 4;13(2):246. doi: 10.3390/v13020246. Viruses. 2021. PMID: 33557422 Free PMC article. Review.
Cited by
-
Structural characterization of a protein adsorbed on aluminum hydroxide adjuvant in vaccine formulation.NPJ Vaccines. 2019 May 28;4:20. doi: 10.1038/s41541-019-0115-7. eCollection 2019. NPJ Vaccines. 2019. PMID: 31149351 Free PMC article.
-
Easy Synthesis of Complex Biomolecular Assemblies: Wheat Germ Cell-Free Protein Expression in Structural Biology.Front Mol Biosci. 2021 Mar 25;8:639587. doi: 10.3389/fmolb.2021.639587. eCollection 2021. Front Mol Biosci. 2021. PMID: 33842544 Free PMC article. Review.
-
Updating on Roles of HIV Intrinsic Factors: A Review of Their Antiviral Mechanisms and Emerging Functions.Intervirology. 2022;65(2):67-79. doi: 10.1159/000519241. Epub 2021 Aug 31. Intervirology. 2022. PMID: 34464956 Free PMC article. Review.
-
Magic-angle-spinning NMR structure of the kinesin-1 motor domain assembled with microtubules reveals the elusive neck linker orientation.Nat Commun. 2022 Nov 10;13(1):6795. doi: 10.1038/s41467-022-34026-w. Nat Commun. 2022. PMID: 36357375 Free PMC article.
-
Mathematical determination of the HIV-1 matrix shell structure and its impact on the biology of HIV-1.PLoS One. 2019 Nov 12;14(11):e0224965. doi: 10.1371/journal.pone.0224965. eCollection 2019. PLoS One. 2019. PMID: 31714942 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

