A Comparison of Techniques for Collecting Skin Microbiome Samples: Swabbing Versus Tape-Stripping
- PMID: 30333815
- PMCID: PMC6176111
- DOI: 10.3389/fmicb.2018.02362
A Comparison of Techniques for Collecting Skin Microbiome Samples: Swabbing Versus Tape-Stripping
Erratum in
-
Corrigendum: A Comparison of Techniques for Collecting Skin Microbiome Samples: Swabbing Versus Tape-Stripping.Front Microbiol. 2018 Nov 20;9:2812. doi: 10.3389/fmicb.2018.02812. eCollection 2018. Front Microbiol. 2018. PMID: 30519225 Free PMC article.
Abstract
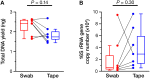
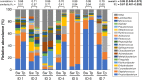
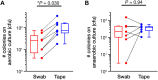
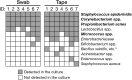
The swabbing and tape-stripping methods have traditionally been used for collecting skin microbiome samples for skin bacterial analysis, although no reports have compared the outcome of these methods for collecting skin bacteria. Our purpose was to show the differences in microbial composition between samples collected using the swabbing and tape-stripping methods, by both the next generation sequencing and culture studies. The skin microbiome was collected by both methods, and the samples were processed for a sequence-based microbiome analysis and culture study. The next-generation sequencing results showed that skin bacteria collected using the tape-stripping method were comparable to those collected using the swabbing method. In the culture study, the tape-stripping method collected a greater number and wider variety of viable skin bacteria than the swabbing method. These results suggest that the tape-stripping method is comparable to the swabbing method for collecting viable skin bacteria, without losing fidelity to the composition of skin microbiome.
Keywords: bacterial culture; next generation sequencing; skin microbiome; swabbing; tape stripping.
Figures


References
-
- Benjamini Y., Hochberg Y. (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B Stat. Methodol. 57 289–300.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

