The Combined Loss of Triose Phosphate and Xylulose 5-Phosphate/Phosphate Translocators Leads to Severe Growth Retardation and Impaired Photosynthesis in Arabidopsis thaliana tpt/xpt Double Mutants
- PMID: 30333839
- PMCID: PMC6175978
- DOI: 10.3389/fpls.2018.01331
The Combined Loss of Triose Phosphate and Xylulose 5-Phosphate/Phosphate Translocators Leads to Severe Growth Retardation and Impaired Photosynthesis in Arabidopsis thaliana tpt/xpt Double Mutants
Abstract
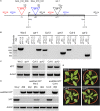
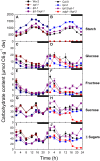
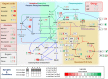
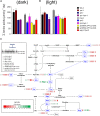
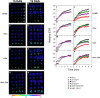
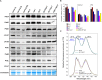
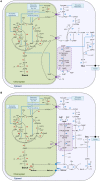
The xylulose 5-phosphate/phosphate translocator (XPT) represents the fourth functional member of the phosphate translocator (PT) family residing in the plastid inner envelope membrane. In contrast to the other three members, little is known on the physiological role of the XPT. Based on its major transport substrates (i.e., pentose phosphates) the XPT has been proposed to act as a link between the plastidial and extraplastidial branches of the oxidative pentose phosphate pathway (OPPP). As the XPT is also capable of transporting triose phosphates, it might as well support the triose phosphate PT (TPT) in exporting photoassimilates from the chloroplast in the light ('day path of carbon') and hence in supplying the whole plant with carbohydrates. Two independent knockout mutant alleles of the XPT (xpt-1 and xpt-2) lacked any specific phenotype, suggesting that the XPT function is redundant. However, double mutants generated from crossings of xpt-1 to different mutant alleles of the TPT (tpt-1 and tpt-2) were severely retarded in size, exhibited a high chlorophyll fluorescence phenotype, and impaired photosynthetic electron transport rates. In the double mutant the export of triose phosphates from the chloroplasts is completely blocked. Hence, precursors for sucrose biosynthesis derive entirely from starch turnover ('night path of carbon'), which was accompanied by a marked accumulation of maltose as a starch breakdown product. Moreover, pentose phosphates produced by the extraplastidial branch of the OPPP also accumulated in the double mutants. Thus, an active XPT indeed retrieves excessive pentose phosphates from the extra-plastidial space and makes them available to the plastids. Further metabolic profiling revealed that phosphorylated intermediates remained largely unaffected, whereas fumarate and glycine contents were diminished in the double mutants. The assessment of C/N-ratios suggested co-limitations of C- and N-metabolism as possible cause for growth retardation of the double mutants. Feeding of sucrose partially rescued the growth and photosynthesis phenotypes of the double mutants. Immunoblots of thylakoid proteins, spectroscopic determinations of photosynthesis complexes, and chlorophyll a fluorescence emission spectra at 77 Kelvin could only partially explain constrains in photosynthesis observed in the double mutants. The data are discussed together with aspects of the OPPP and central carbon metabolism.
Keywords: chloroplasts; pentose phosphates; photosynthesis; signaling; transport.
Figures


Similar articles
-
The Xylulose 5-Phosphate/Phosphate Translocator Supports Triose Phosphate, but Not Phosphoenolpyruvate Transport Across the Inner Envelope Membrane of Plastids in Arabidopsis thaliana Mutant Plants.Front Plant Sci. 2018 Oct 18;9:1461. doi: 10.3389/fpls.2018.01461. eCollection 2018. Front Plant Sci. 2018. PMID: 30405650 Free PMC article.
-
Defects in leaf carbohydrate metabolism compromise acclimation to high light and lead to a high chlorophyll fluorescence phenotype in Arabidopsis thaliana.BMC Plant Biol. 2012 Jan 16;12:8. doi: 10.1186/1471-2229-12-8. BMC Plant Biol. 2012. PMID: 22248311 Free PMC article.
-
An Arabidopsis thaliana knock-out mutant of the chloroplast triose phosphate/phosphate translocator is severely compromised only when starch synthesis, but not starch mobilisation is abolished.Plant J. 2002 Dec;32(5):685-99. doi: 10.1046/j.1365-313x.2002.01460.x. Plant J. 2002. PMID: 12472685
-
Solute transporters as connecting elements between cytosol and plastid stroma.Curr Opin Plant Biol. 2004 Jun;7(3):247-53. doi: 10.1016/j.pbi.2004.03.008. Curr Opin Plant Biol. 2004. PMID: 15134744 Review.
-
Transfer of substrates across the chloroplast envelope.Horiz Biochem Biophys. 1976;2:199-299. Horiz Biochem Biophys. 1976. PMID: 776769 Review.
Cited by
-
The Arabidopsis Plastidial Glucose-6-Phosphate Transporter GPT1 is Dually Targeted to Peroxisomes via the Endoplasmic Reticulum.Plant Cell. 2020 May;32(5):1703-1726. doi: 10.1105/tpc.19.00959. Epub 2020 Feb 28. Plant Cell. 2020. PMID: 32111666 Free PMC article.
-
The plastidial pentose phosphate pathway is essential for postglobular embryo development in Arabidopsis.Proc Natl Acad Sci U S A. 2019 Jul 23;116(30):15297-15306. doi: 10.1073/pnas.1908556116. Epub 2019 Jul 11. Proc Natl Acad Sci U S A. 2019. PMID: 31296566 Free PMC article.
-
BnaPPT1 is essential for chloroplast development and seed oil accumulation in Brassica napus.J Adv Res. 2022 Dec;42:29-40. doi: 10.1016/j.jare.2022.07.008. Epub 2022 Jul 27. J Adv Res. 2022. PMID: 35907629 Free PMC article.
-
From source to sink: mechanistic insight of photoassimilates synthesis and partitioning under high temperature and elevated [CO2].Plant Mol Biol. 2022 Nov;110(4-5):305-324. doi: 10.1007/s11103-022-01274-9. Epub 2022 May 24. Plant Mol Biol. 2022. PMID: 35610527 Review.
-
Transcriptional Regulation of the Glucose-6-Phosphate/Phosphate Translocator 2 Is Related to Carbon Exchange Across the Chloroplast Envelope.Front Plant Sci. 2019 Jun 27;10:827. doi: 10.3389/fpls.2019.00827. eCollection 2019. Front Plant Sci. 2019. PMID: 31316533 Free PMC article.
References
-
- Arrivault S., Guenther M., Ivakov A., Feil R., Vosloh D., van Dongen J. T., et al. (2009). Use of reverse-phase liquid chromatography, linked to tandem mass spectrometry, to profile the Calvin cycle and other metabolic intermediates in Arabidopsis rosettes at different carbon dioxide concentrations. Plant J. 59 824–839. 10.1111/j.1365-313X.2009.03902.x - DOI - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

