A RNA producing DNA hydrogel as a platform for a high performance RNA interference system
- PMID: 30337586
- PMCID: PMC6193956
- DOI: 10.1038/s41467-018-06864-0
A RNA producing DNA hydrogel as a platform for a high performance RNA interference system
Abstract
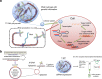
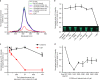
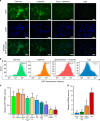
RNA interference (RNAi) is a mechanism in which small interfering RNA (siRNA) silences a target gene. Herein, we describe a DNA hydrogel capable of producing siRNA and interfering with protein expression. This RNAi-exhibiting gel (termed I-gel for interfering gel) consists of a plasmid carrying the gene transcribing siRNA against the target mRNA as part of the gel scaffold. The RNAi efficiency of the I-gel has been confirmed by green fluorescent protein (GFP) expression assay and RNA production quantification. The plasmid stability in the I-gel results in an 8-times higher transcription efficiency than that of the free plasmid. We further applied the I-gel to live cells and confirmed its effect in interfering with the GFP expression. The I-gel shows higher RNAi effect than plasmids in free form or complexed with Lipofectamine. This nanoscale hydrogel, which is able to produce RNA in a cell, provides a platform technology for efficient RNAi system.
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
[Detection of RNA interference in nasopharyngeal carcinoma cell lines using reporter genes].Ai Zheng. 2005 Mar;24(3):371-5. Ai Zheng. 2005. PMID: 15757546 Chinese.
-
Application of RNA interference to chicken embryos using small interfering RNA.J Exp Zool A Comp Exp Biol. 2004 Oct 1;301(10):820-7. doi: 10.1002/jez.a.99. J Exp Zool A Comp Exp Biol. 2004. PMID: 15449340
-
Double Controlled Release of Therapeutic RNA Modules through Injectable DNA-RNA Hybrid Hydrogel.ACS Appl Mater Interfaces. 2020 Dec 16;12(50):55554-55563. doi: 10.1021/acsami.0c12506. Epub 2020 Dec 1. ACS Appl Mater Interfaces. 2020. PMID: 33259200
-
Engineered Hydrogels for Local and Sustained Delivery of RNA-Interference Therapies.Adv Healthc Mater. 2017 Jan;6(1):10.1002/adhm.201601041. doi: 10.1002/adhm.201601041. Epub 2016 Dec 15. Adv Healthc Mater. 2017. PMID: 27976524 Free PMC article. Review.
-
Nonviral vector-mediated RNA interference: its gene silencing characteristics and important factors to achieve RNAi-based gene therapy.Adv Drug Deliv Rev. 2009 Jul 25;61(9):760-6. doi: 10.1016/j.addr.2009.04.006. Epub 2009 Apr 20. Adv Drug Deliv Rev. 2009. PMID: 19386274 Review.
Cited by
-
Hydrogels as functional components in artificial cell systems.Nat Rev Chem. 2022 Aug;6(8):562-578. doi: 10.1038/s41570-022-00404-7. Epub 2022 Jul 27. Nat Rev Chem. 2022. PMID: 37118012 Review.
-
Dynamic Covalent Hydrogels: Strong yet Dynamic.Gels. 2022 Sep 10;8(9):577. doi: 10.3390/gels8090577. Gels. 2022. PMID: 36135289 Free PMC article. Review.
-
Stimuli-Responsive DNA Hydrogel Design Strategies for Biomedical Applications.Biosensors (Basel). 2025 Jun 4;15(6):355. doi: 10.3390/bios15060355. Biosensors (Basel). 2025. PMID: 40558437 Free PMC article. Review.
-
Smart Bioinks as de novo Building Blocks to Bioengineer Living Tissues.Gels. 2019 May 22;5(2):29. doi: 10.3390/gels5020029. Gels. 2019. PMID: 31121889 Free PMC article.
-
pH-Sensitive Chitosan-Heparin Nanoparticles for Effective Delivery of Genetic Drugs into Epithelial Cells.Pharmaceutics. 2019 Jul 5;11(7):317. doi: 10.3390/pharmaceutics11070317. Pharmaceutics. 2019. PMID: 31284414 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

