In vivo RNA structural probing of uracil and guanine base-pairing by 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC)
- PMID: 30341176
- PMCID: PMC6298566
- DOI: 10.1261/rna.067868.118
In vivo RNA structural probing of uracil and guanine base-pairing by 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC)
Abstract
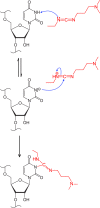
Many biological functions performed by RNAs arise from their in vivo structures. The structure of the same RNA can differ in vitro and in vivo owing in part to the influence of molecules ranging from protons to secondary metabolites to proteins. Chemical reagents that modify the Watson-Crick (WC) face of unprotected RNA bases report on the absence of base-pairing and so are of value to determining structures adopted by RNAs. Reagents have thus been sought that can report on the native RNA structures that prevail in living cells. Dimethyl sulfate (DMS) and glyoxal penetrate cell membranes and inform on RNA secondary structure in vivo through modification of adenine (A), cytosine (C), and guanine (G) bases. Uracil (U) bases, however, have thus far eluded characterization in vivo. Herein, we show that the water-soluble carbodiimide 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) is capable of modifying the WC face of U and G in vivo, favoring the former nucleobase by a factor of ∼1.5, and doing so in the eukaryote rice, as well as in the Gram-negative bacterium Escherichia coli While both EDC and glyoxal target Gs, EDC reacts with Gs in their typical neutral state, while glyoxal requires Gs to populate the rare anionic state. EDC may thus be more generally useful; however, comparison of the reactivity of EDC and glyoxal may allow the identification of Gs with perturbed pKas in vivo and genome-wide. Overall, use of EDC with DMS allows in vivo probing of the base-pairing status of all four RNA bases.
Keywords: 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide; EDC; RNA structure; in vivo RNA probing.
© 2019 Mitchell et al.; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures





Similar articles
-
Carbodiimide reagents for the chemical probing of RNA structure in cells.RNA. 2019 Jan;25(1):135-146. doi: 10.1261/rna.067561.118. Epub 2018 Nov 2. RNA. 2019. PMID: 30389828 Free PMC article.
-
A new reagent for in vivo structure probing of RNA G and U residues that improves RNA structure prediction alone and combined with DMS.RNA. 2024 Jun 17;30(7):901-919. doi: 10.1261/rna.079974.124. RNA. 2024. PMID: 38670632 Free PMC article.
-
Glyoxals as in vivo RNA structural probes of guanine base-pairing.RNA. 2018 Jan;24(1):114-124. doi: 10.1261/rna.064014.117. Epub 2017 Oct 13. RNA. 2018. PMID: 29030489 Free PMC article.
-
Probing RNA structure in vivo.Curr Opin Struct Biol. 2019 Dec;59:151-158. doi: 10.1016/j.sbi.2019.07.008. Epub 2019 Sep 13. Curr Opin Struct Biol. 2019. PMID: 31521910 Free PMC article. Review.
-
The G x U wobble base pair. A fundamental building block of RNA structure crucial to RNA function in diverse biological systems.EMBO Rep. 2000 Jul;1(1):18-23. doi: 10.1093/embo-reports/kvd001. EMBO Rep. 2000. PMID: 11256617 Free PMC article. Review.
Cited by
-
Viral RNA structure analysis using DMS-MaPseq.Methods. 2020 Nov 1;183:68-75. doi: 10.1016/j.ymeth.2020.04.001. Epub 2020 Apr 3. Methods. 2020. PMID: 32251733 Free PMC article. Review.
-
Mutation signature filtering enables high-fidelity RNA structure probing at all four nucleobases with DMS.Nucleic Acids Res. 2023 Sep 8;51(16):8744-8757. doi: 10.1093/nar/gkad522. Nucleic Acids Res. 2023. PMID: 37334863 Free PMC article.
-
Rock, scissors, paper: How RNA structure informs function.Plant Cell. 2023 May 29;35(6):1671-1707. doi: 10.1093/plcell/koad026. Plant Cell. 2023. PMID: 36747354 Free PMC article.
-
Tissue-specific changes in the RNA structurome mediate salinity response in Arabidopsis.RNA. 2020 Apr;26(4):492-511. doi: 10.1261/rna.072850.119. Epub 2020 Jan 14. RNA. 2020. PMID: 31937672 Free PMC article.
-
In vivo structure probing of RNA in Archaea: novel insights into the ribosome structure of Methanosarcina acetivorans.RNA. 2023 Oct;29(10):1610-1620. doi: 10.1261/rna.079687.123. Epub 2023 Jul 25. RNA. 2023. PMID: 37491319 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
