A multi-tissue full lifespan epigenetic clock for mice
- PMID: 30348905
- PMCID: PMC6224226
- DOI: 10.18632/aging.101590
A multi-tissue full lifespan epigenetic clock for mice
Abstract
Human DNA-methylation data have been used to develop highly accurate biomarkers of aging ("epigenetic clocks"). Recent studies demonstrate that similar epigenetic clocks for mice (Mus Musculus) can be slowed by gold standard anti-aging interventions such as calorie restriction and growth hormone receptor knock-outs. Using DNA methylation data from previous publications with data collected in house for a total 1189 samples spanning 193,651 CpG sites, we developed 4 novel epigenetic clocks by choosing different regression models (elastic net- versus ridge regression) and by considering different sets of CpGs (all CpGs vs highly conserved CpGs). We demonstrate that accurate age estimators can be built on the basis of highly conserved CpGs. However, the most accurate clock results from applying elastic net regression to all CpGs. While the anti-aging effect of calorie restriction could be detected with all types of epigenetic clocks, only ridge regression based clocks replicated the finding of slow epigenetic aging effects in dwarf mice. Overall, this study demonstrates that there are trade-offs when it comes to epigenetic clocks in mice. Highly accurate clocks might not be optimal for detecting the beneficial effects of anti-aging interventions.
Keywords: DNA methylation; biological age; epigenetic clock; mouse.
Conflict of interest statement
Figures

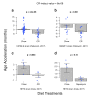
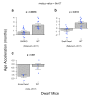


Similar articles
-
Epigenetic aging signatures in mice livers are slowed by dwarfism, calorie restriction and rapamycin treatment.Genome Biol. 2017 Mar 28;18(1):57. doi: 10.1186/s13059-017-1186-2. Genome Biol. 2017. PMID: 28351423 Free PMC article.
-
Hepatic gene body hypermethylation is a shared epigenetic signature of murine longevity.PLoS Genet. 2018 Nov 21;14(11):e1007766. doi: 10.1371/journal.pgen.1007766. eCollection 2018 Nov. PLoS Genet. 2018. PMID: 30462643 Free PMC article.
-
Many chronological aging clocks can be found throughout the epigenome: Implications for quantifying biological aging.Aging Cell. 2021 Nov;20(11):e13492. doi: 10.1111/acel.13492. Epub 2021 Oct 16. Aging Cell. 2021. PMID: 34655509 Free PMC article.
-
DNA Methylation Clocks in Aging: Categories, Causes, and Consequences.Mol Cell. 2018 Sep 20;71(6):882-895. doi: 10.1016/j.molcel.2018.08.008. Mol Cell. 2018. PMID: 30241605 Free PMC article. Review.
-
Analysis of epigenetic aging in vivo and in vitro: Factors controlling the speed and direction.Exp Biol Med (Maywood). 2020 Nov;245(17):1543-1551. doi: 10.1177/1535370220947015. Epub 2020 Aug 6. Exp Biol Med (Maywood). 2020. PMID: 32762265 Free PMC article. Review.
Cited by
-
Profiling epigenetic age in single cells.Nat Aging. 2021 Dec;1(12):1189-1201. doi: 10.1038/s43587-021-00134-3. Epub 2021 Dec 9. Nat Aging. 2021. PMID: 36211119 Free PMC article.
-
The immunity and redox clocks in mice, markers of lifespan.Sci Rep. 2024 Jan 19;14(1):1703. doi: 10.1038/s41598-024-51978-9. Sci Rep. 2024. PMID: 38242936 Free PMC article.
-
The Genetics of Aging: A Vertebrate Perspective.Cell. 2019 Mar 21;177(1):200-220. doi: 10.1016/j.cell.2019.02.038. Cell. 2019. PMID: 30901541 Free PMC article. Review.
-
The relationship between epigenetic age and the hallmarks of aging in human cells.Nat Aging. 2022 Jun;2(6):484-493. doi: 10.1038/s43587-022-00220-0. Epub 2022 May 16. Nat Aging. 2022. PMID: 37034474 Free PMC article.
-
The effects of age, sex, weight, and breed on canid methylomes.Epigenetics. 2022 Nov;17(11):1497-1512. doi: 10.1080/15592294.2022.2069385. Epub 2022 May 3. Epigenetics. 2022. PMID: 35502722 Free PMC article.
References
-
- Christensen BC, Houseman EA, Marsit CJ, Zheng S, Wrensch MR, Wiemels JL, Nelson HH, Karagas MR, Padbury JF, Bueno R, Sugarbaker DJ, Yeh RF, Wiencke JK, Kelsey KT. Aging and environmental exposures alter tissue-specific DNA methylation dependent upon CpG island context. PLoS Genet. 2009; 5:e1000602. 10.1371/journal.pgen.1000602 - DOI - PMC - PubMed
-
- Rakyan VK, Down TA, Maslau S, Andrew T, Yang TP, Beyan H, Whittaker P, McCann OT, Finer S, Valdes AM, Leslie RD, Deloukas P, Spector TD. Human aging-associated DNA hypermethylation occurs preferentially at bivalent chromatin domains. Genome Res. 2010; 20:434–39. 10.1101/gr.103101.109 - DOI - PMC - PubMed
-
- Teschendorff AE, Menon U, Gentry-Maharaj A, Ramus SJ, Weisenberger DJ, Shen H, Campan M, Noushmehr H, Bell CG, Maxwell AP, Savage DA, Mueller-Holzner E, Marth C, et al.. Age-dependent DNA methylation of genes that are suppressed in stem cells is a hallmark of cancer. Genome Res. 2010; 20:440–46. 10.1101/gr.103606.109 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

