Understanding Conformational Dynamics of Complex Lipid Mixtures Relevant to Biology
- PMID: 30350011
- PMCID: PMC6244758
- DOI: 10.1007/s00232-018-0050-y
Understanding Conformational Dynamics of Complex Lipid Mixtures Relevant to Biology
Abstract
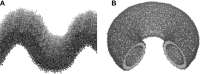
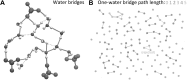
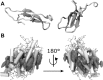
This is a perspective article entitled "Frontiers in computational biophysics: understanding conformational dynamics of complex lipid mixtures relevant to biology" which is following a CECAM meeting with the same name.
Keywords: Cell membrane; Computational biophysics; Lipid–protein interactions; Molecular dynamics.
Conflict of interest statement
Conflicts of interest
The authors declare no conflict of interest.
Research Involving Human Participants or Animals
This research does not involve human participants or animals.
Figures




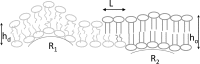





References
-
- Beevers AJ, Kukol A. The transmembrane domain of the oncogenic mutant erbb-2 receptor: a structure obtained from site-specific infrared dichroism and molecular dynamics. J Mol Biol. 2006;361:945–953. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

