Off-axis rotor in Enterococcus hirae V-ATPase visualized by Zernike phase plate single-particle cryo-electron microscopy
- PMID: 30353110
- PMCID: PMC6199243
- DOI: 10.1038/s41598-018-33977-9
Off-axis rotor in Enterococcus hirae V-ATPase visualized by Zernike phase plate single-particle cryo-electron microscopy
Abstract
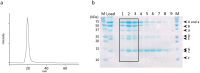
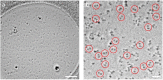
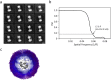
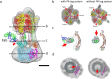
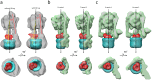
EhV-ATPase is an ATP-driven Na+ pump in the eubacteria Enterococcus hirae (Eh). Here, we present the first entire structure of detergent-solubilized EhV-ATPase by single-particle cryo-electron microscopy (cryo-EM) using Zernike phase plate. The cryo-EM map dominantly showed one of three catalytic conformations in this rotary enzyme. To further stabilize the originally heterogeneous structure caused by the ATP hydrolysis states of the V1-ATPases, a peptide epitope tag system was adopted, in which the inserted peptide epitope sequence interfered with rotation of the central rotor by binding the Fab. As a result, the map unexpectedly showed another catalytic conformation of EhV-ATPase. Interestingly, these two conformations identified with and without Fab conversely coincided with those of the minor state 2 and the major state 1 of Thermus thermophilus V/A-ATPase, respectively. The most prominent feature in EhV-ATPase was the off-axis rotor, where the cytoplasmic V1 domain was connected to the transmembrane Vo domain through the off-axis central rotor. Furthermore, compared to the structure of ATP synthases, the larger size of the interface between the transmembrane a-subunit and c-ring of EhV-ATPase would be more advantageous for active ion pumping.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Six states of Enterococcus hirae V-type ATPase reveals non-uniform rotor rotation during turnover.Commun Biol. 2023 Jul 28;6(1):755. doi: 10.1038/s42003-023-05110-8. Commun Biol. 2023. PMID: 37507515 Free PMC article.
-
Cryo EM structure of intact rotary H+-ATPase/synthase from Thermus thermophilus.Nat Commun. 2018 Jan 8;9(1):89. doi: 10.1038/s41467-017-02553-6. Nat Commun. 2018. PMID: 29311594 Free PMC article.
-
Torque generation of Enterococcus hirae V-ATPase.J Biol Chem. 2014 Nov 7;289(45):31212-23. doi: 10.1074/jbc.M114.598177. Epub 2014 Sep 25. J Biol Chem. 2014. PMID: 25258315 Free PMC article.
-
Molecular structure and rotary dynamics of Enterococcus hirae V₁-ATPase.IUBMB Life. 2014 Sep;66(9):624-30. doi: 10.1002/iub.1311. Epub 2014 Sep 17. IUBMB Life. 2014. PMID: 25229752 Review.
-
The study of vacuolar-type ATPases by single particle electron microscopy.Biochem Cell Biol. 2014 Dec;92(6):460-6. doi: 10.1139/bcb-2014-0086. Epub 2014 Sep 3. Biochem Cell Biol. 2014. PMID: 25350640 Review.
Cited by
-
Single-molecule analysis reveals rotational substeps and chemo-mechanical coupling scheme of Enterococcus hirae V1-ATPase.J Biol Chem. 2019 Nov 8;294(45):17017-17030. doi: 10.1074/jbc.RA119.008947. Epub 2019 Sep 13. J Biol Chem. 2019. PMID: 31519751 Free PMC article.
-
Advances in domain and subunit localization technology for electron microscopy.Biophys Rev. 2019 Apr;11(2):149-155. doi: 10.1007/s12551-019-00513-6. Epub 2019 Mar 5. Biophys Rev. 2019. PMID: 30834502 Free PMC article. Review.
-
Use of phase plate cryo-EM reveals conformation diversity of therapeutic IgG with 50 kDa Fab fragment resolved below 6 Å.Sci Rep. 2024 Jun 18;14(1):14079. doi: 10.1038/s41598-024-62045-8. Sci Rep. 2024. PMID: 38890341 Free PMC article.
-
Biophysical research in Okazaki, Japan.Biophys Rev. 2020 Apr;12(2):237-243. doi: 10.1007/s12551-020-00633-4. Epub 2020 Feb 15. Biophys Rev. 2020. PMID: 32062838 Free PMC article. Review.
-
ATP synthesis of Enterococcus hirae V-ATPase driven by sodium motive force.J Biol Chem. 2025 Apr;301(4):108422. doi: 10.1016/j.jbc.2025.108422. Epub 2025 Mar 19. J Biol Chem. 2025. PMID: 40118453 Free PMC article.
References
-
- Yoshimori T, Yamamoto A, Moriyama Y, Futai M, Tashiro Y. Bafilomycin A1, a specific inhibitor of vacuolar-type H+-ATPase, inhibits acidification and protein-degradation in lysosomes of cultured-cells. J. Biol. Chem. 1991;266:17707–17712. - PubMed
-
- Wieczorek H, Brown D, Grinstein S, Ehrenfeld J, Harvey WR. Animal plasma membrane energization by proton-motive V-ATPases. Bio Essays. 1999;21:637–648. - PubMed
-
- Harvey WR. Physiology of V-ATPases. J. Exp. Biol. 1992;172:1–17. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

