Pushing the limits of whole genome amplification: successful sequencing of RADseq library from a single microhymenopteran (Chalcidoidea, Trichogramma)
- PMID: 30356952
- PMCID: PMC6195110
- DOI: 10.7717/peerj.5640
Pushing the limits of whole genome amplification: successful sequencing of RADseq library from a single microhymenopteran (Chalcidoidea, Trichogramma)
Abstract
A major obstacle to high-throughput genotyping of microhymenoptera is their small size. As species are difficult to discriminate, and because complexes may exist, the sequencing of a pool of specimens is hazardous. Thus, one should be able to sequence pangenomic markers (e.g., RADtags) from a single specimen. To date, whole genome amplification (WGA) prior to library construction is still a necessity as at most 10 ng of DNA can be obtained from single specimens (sometimes less). However, this amount of DNA is not compatible with manufacturer's requirements for commercial kits. Here we test the accuracy of the GenomiPhi kit V2 on Trichogramma wasps by comparing RAD libraries obtained from the WGA of single specimens (F0 and F1 generation, about1 ng input DNA for the WGA (0.17-2.9 ng)) and a biological amplification of genomic material (the pool of the progeny of the F1 generation). Globally, we found that 99% of the examined loci (up to 48,189 for one of the crosses, 109 bp each) were compatible with the mode of reproduction of the studied model (haplodiploidy) and Mendelian inheritance of alleles. The remaining 1% (0.01% of the analysed nucleotides) could represent WGA bias or other experimental/analytical bias. This study shows that the multiple displacement amplification method on which the GenomiPhi kit relies, could also be of great help for the high-throughput genotyping of microhymenoptera used for biological control, or other organisms from which only a very small amount of DNA can be extracted, such as human disease vectors (e.g., sandflies, fleas, ticks etc.).
Keywords: GenomiPhi; High-throughput genotyping; Microarthropods; RAD; Small amount of DNA; WGA.
Conflict of interest statement
The authors declare there are no competing interests.
Figures
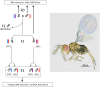
References
-
- Al Khatib F, Fusu L, Cruaud A, Gibson G, Borowiec N, Ris N, Rasplus J-Y, Delvare G. An integrative approach for species discrimination in the Eupelmus urozonus complex (Hymenoptera, Eupelmidae), with the descriptions of eleven new species from the West Palaearctic. Systematic Entomology. 2014;39:806–862. doi: 10.1111/syen.12089. - DOI
-
- Austin AD, Dowton M, editors. Hymenoptera: evolution, biodiversity and biological control. Collingwood: CSIRO PUBLISHING; 2000.
-
- Barker DL, Hansen MS, Faruqi AF, Giannola D, Irsula OR, Lasken RS, Latterich M, Makarov V, Oliphant A, Pinter JH, Shen R, Sleptsova I, Ziehler W, Lai E. Two methods of whole-genome amplification enable accurate genotyping across a 2320-SNP linkage panel. Genome Research. 2004;14:901–907. doi: 10.1101/gr.1949704. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

