Cerebral Cavernous Malformations Develop Through Clonal Expansion of Mutant Endothelial Cells
- PMID: 30359189
- PMCID: PMC6205520
- DOI: 10.1161/CIRCRESAHA.118.313970
Cerebral Cavernous Malformations Develop Through Clonal Expansion of Mutant Endothelial Cells
Abstract
Rationale: Vascular malformations arise in vessels throughout the entire body. Causative genetic mutations have been identified for many of these diseases; however, little is known about the mutant cell lineage within these malformations.
Objective: We utilize an inducible mouse model of cerebral cavernous malformations (CCMs) coupled with a multicolor fluorescent reporter to visualize the contribution of mutant endothelial cells (ECs) to the malformation.
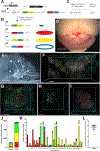
Methods and results: We combined a Ccm3 mouse model with the confetti fluorescent reporter to simultaneously delete Ccm3 and label the mutant EC with 1 of 4 possible colors. We acquired Z-series confocal images from serial brain sections and created 3-dimensional reconstructions of entire CCMs to visualize mutant ECs during CCM development. We observed a pronounced pattern of CCMs lined with mutant ECs labeled with a single confetti color (n=42). The close 3-dimensional distribution, as determined by the nearest neighbor analysis, of the clonally dominant ECs within the CCM was statistically different than the background confetti labeling of ECs in non-CCM control brain slices as well as a computer simulation ( P<0.001). Many of the small (<100 μm diameter) CCMs consisted, almost exclusively, of the clonally dominant mutant ECs labeled with the same confetti color, whereas the large (>100 μm diameter) CCMs contained both the clonally dominant mutant cells and wild-type ECs. We propose of model of CCM development in which an EC acquires a second somatic mutation, undergoes clonal expansion to initiate CCM formation, and then incorporates neighboring wild-type ECs to increase the size of the malformation.
Conclusions: This is the first study to visualize, with single-cell resolution, the clonal expansion of mutant ECs within CCMs. The incorporation of wild-type ECs into the growing malformation presents another series of cellular events whose elucidation would enhance our understanding of CCMs and may provide novel therapeutic opportunities.
Keywords: cell lineage; cerebral cavernous malformations; cerebrovascular disease; genetics; mutation; vascular malformation.
Figures



References
-
- Queisser A, Boon LM and Vikkula M. Etiology and Genetics of Congenital Vascular Lesions. Otolaryngologic clinics of North America. 2018;51:41–53. - PubMed
-
- Sahoo T, Johnson EW, Thomas JW, Kuehl PM, Jones TL, Dokken CG, Touchman JW, Gallione CJ, Lee-Lin SQ, Kosofsky B, Kurth JH, Louis DN, Mettler G, Morrison L, Gil-Nagel A, Rich SS, Zabramski JM, Boguski MS, Green ED and Marchuk DA. Mutations in the gene encoding KRIT1, a Krev-1/rap1a binding protein, cause cerebral cavernous malformations (CCM1). Human molecular genetics. 1999;8:2325–33. - PubMed
-
- Laberge-le Couteulx S, Jung HH, Labauge P, Houtteville JP, Lescoat C, Cecillon M, Marechal E, Joutel A, Bach JF and Tournier-Lasserve E. Truncating mutations in CCM1, encoding KRIT1, cause hereditary cavernous angiomas. Nature genetics. 1999;23:189–93. - PubMed
-
- Liquori CL, Berg MJ, Siegel AM, Huang E, Zawistowski JS, Stoffer T, Verlaan D, Balogun F, Hughes L, Leedom TP, Plummer NW, Cannella M, Maglione V, Squitieri F, Johnson EW, Rouleau GA, Ptacek L and Marchuk DA. Mutations in a gene encoding a novel protein containing a phosphotyrosine-binding domain cause type 2 cerebral cavernous malformations. American journal of human genetics. 2003;73:1459–64. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

