A single amino acid distorts the Fc γ receptor IIIb/CD16b structure upon binding immunoglobulin G1 and reduces affinity relative to CD16a
- PMID: 30361439
- PMCID: PMC6314127
- DOI: 10.1074/jbc.RA118.005273
A single amino acid distorts the Fc γ receptor IIIb/CD16b structure upon binding immunoglobulin G1 and reduces affinity relative to CD16a
Abstract
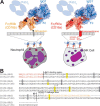
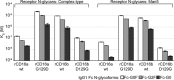
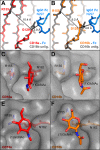
Therapeutic mAbs engage Fc γ receptor III (CD16) to elicit a protective cell-mediated response and destroy the target tissue. Newer drugs designed to bind CD16a with increased affinity surprisingly also elicit protective CD16b-mediated responses. However, it is unclear why IgG binds CD16a with more than 10-fold higher affinity than CD16b even though these receptors share more than 97% identity. Here we identified one residue, Gly-129, that contributes to the greater IgG binding affinity of CD16a. The CD16b variant D129G bound IgG1 Fc with 2-fold higher affinity than CD16a and with 90-fold higher affinity than the WT. Conversely, the binding affinity of CD16a-G129D was decreased 128-fold relative to WT CD16a and comparably to that of WT CD16b. The interaction of IgG1 Fc with CD16a, but not with CD16b, is known to be sensitive to the composition of the asparagine-linked carbohydrates (N-glycans) attached to the receptor. CD16a and CD16b-D129G displaying minimally processed oligomannose N-glycans bound to IgG1 Fc with about 5.2-fold increased affinity compared with variants with highly processed complex-type N-glycans. CD16b and the CD16a-G129D variant exhibited a smaller 1.9-fold affinity increase with oligomannose N-glycans. A model of glycosylated CD16b bound to IgG1 Fc determined to 2.2 Å resolution combined with a 250-ns all-atom molecular dynamics simulation showed that the larger Asp-129 residue deformed the Fc-binding surface. These results reveal how Asp-129 in CD16b affects its binding affinity for IgG1 Fc and suggest that antibodies engineered to engage CD16b with high affinity must accommodate the Asp-129 side chain.
Keywords: IgG; N-linked glycosylation; X-ray crystallography; fragment crystallizable region; immunity; molecular dynamics; monoclonal antibody; posttranslational modification (PTM); protein–protein interaction.
© 2018 Roberts and Barb.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the Department of Energy
Figures




References
-
- Hayes J. M., Cosgrave E. F., Struwe W. B., Wormald M., Davey G. P., Jefferis R., and Rudd P. M. (2014) Glycosylation and Fc receptors. Curr. Top. Microbiol. Immunol. 382, 165–199 - PubMed
-
- Dekkers G., Treffers L., Plomp R., Bentlage A. E. H., de Boer M., Koeleman C. A. M., Lissenberg-Thunnissen S. N., Visser R., Brouwer M., Mok J. Y., Matlung H., van den Berg T. K., van Esch W. J. E., Kuijpers T. W., Wouters D., et al. (2017) Decoding the human immunoglobulin G-glycan repertoire reveals a spectrum of Fc receptor- and complement-mediated effector activities. Front. Immunol. 8, 877 10.3389/fimmu.2017.00877 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

