Structural dynamics of the E6AP/UBE3A-E6-p53 enzyme-substrate complex
- PMID: 30361475
- PMCID: PMC6202321
- DOI: 10.1038/s41467-018-06953-0
Structural dynamics of the E6AP/UBE3A-E6-p53 enzyme-substrate complex
Abstract
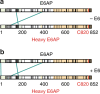
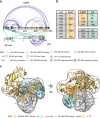
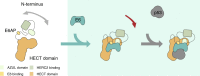
Deregulation of the ubiquitin ligase E6AP is causally linked to the development of human disease, including cervical cancer. In complex with the E6 oncoprotein of human papillomaviruses, E6AP targets the tumor suppressor p53 for degradation, thereby contributing to carcinogenesis. Moreover, E6 acts as a potent activator of E6AP by a yet unknown mechanism. However, structural information explaining how the E6AP-E6-p53 enzyme-substrate complex is assembled, and how E6 stimulates E6AP, is largely missing. Here, we develop and apply different crosslinking mass spectrometry-based approaches to study the E6AP-E6-p53 interplay. We show that binding of E6 induces conformational rearrangements in E6AP, thereby positioning E6 and p53 in the immediate vicinity of the catalytic center of E6AP. Our data provide structural and functional insights into the dynamics of the full-length E6AP-E6-p53 enzyme-substrate complex, demonstrating how E6 can stimulate the ubiquitin ligase activity of E6AP while facilitating ubiquitin transfer from E6AP onto p53.
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Structure of E6AP in complex with HPV16-E6 and p53 reveals a novel ordered domain important for E3 ligase activation.Structure. 2025 Mar 6;33(3):504-516.e4. doi: 10.1016/j.str.2024.12.013. Epub 2025 Jan 15. Structure. 2025. PMID: 39818213
-
Stepwise multipolyubiquitination of p53 by the E6AP-E6 ubiquitin ligase complex.J Biol Chem. 2019 Oct 11;294(41):14860-14875. doi: 10.1074/jbc.RA119.008374. Epub 2019 Sep 6. J Biol Chem. 2019. PMID: 31492752 Free PMC article.
-
Structure of High-Risk Papillomavirus 31 E6 Oncogenic Protein and Characterization of E6/E6AP/p53 Complex Formation.J Virol. 2020 Dec 22;95(2):e00730-20. doi: 10.1128/JVI.00730-20. Print 2020 Dec 22. J Virol. 2020. PMID: 33115863 Free PMC article.
-
HPV E6, E6AP and cervical cancer.BMC Biochem. 2008 Oct 21;9 Suppl 1(Suppl 1):S4. doi: 10.1186/1471-2091-9-S1-S4. BMC Biochem. 2008. PMID: 19007434 Free PMC article. Review.
-
The role of TP53 in Cervical carcinogenesis.Hum Mutat. 2003 Mar;21(3):307-12. doi: 10.1002/humu.10178. Hum Mutat. 2003. PMID: 12619117 Review.
Cited by
-
Kinetic principles of chemical cross-link formation for protein-protein interactions.Proc Natl Acad Sci U S A. 2024 Dec 17;121(51):e2402040121. doi: 10.1073/pnas.2402040121. Epub 2024 Dec 9. Proc Natl Acad Sci U S A. 2024. PMID: 39652756 Free PMC article.
-
Structure of E6AP in complex with HPV16-E6 and p53 reveals a novel ordered domain important for E3 ligase activation.Structure. 2025 Mar 6;33(3):504-516.e4. doi: 10.1016/j.str.2024.12.013. Epub 2025 Jan 15. Structure. 2025. PMID: 39818213
-
Optimized isolation of enzymatically active ubiquitin E3 ligase E6AP/UBE3A from mammalian cells.Protein Expr Purif. 2025 Apr;228:106661. doi: 10.1016/j.pep.2025.106661. Epub 2025 Jan 9. Protein Expr Purif. 2025. PMID: 39798888
-
Potential role of human papillomavirus proteins associated with the development of cancer.Virusdisease. 2022 Sep;33(3):322-333. doi: 10.1007/s13337-022-00786-8. Epub 2022 Aug 27. Virusdisease. 2022. PMID: 36277412 Free PMC article. Review.
-
A ubiquitin variant-based affinity approach selectively identifies substrates of the ubiquitin ligase E6AP in complex with HPV-11 E6 or HPV-16 E6.J Biol Chem. 2020 Oct 30;295(44):15070-15082. doi: 10.1074/jbc.RA120.015603. Epub 2020 Aug 27. J Biol Chem. 2020. PMID: 32855237 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
- SFB 969/Deutsche Forschungsgemeinschaft (German Research Foundation)/International
- BB/N015274/1/Biotechnology and Biological Sciences Research Council (BBSRC)/International
- Wellcome Trust/United Kingdom
- STE 2517/1-1/Deutsche Forschungsgemeinschaft (German Research Foundation)/International
- 104194/Z/14/Z/Wellcome Trust/United Kingdom
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

