Long-term positioning and polar preference of chemoreceptor clusters in E. coli
- PMID: 30361683
- PMCID: PMC6202326
- DOI: 10.1038/s41467-018-06835-5
Long-term positioning and polar preference of chemoreceptor clusters in E. coli
Abstract
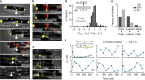
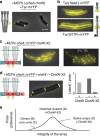
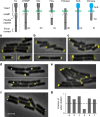
The bacterial chemosensory arrays are a notable model for studying the basic principles of receptor clustering and cellular organization. Here, we provide a new perspective regarding the long-term dynamics of these clusters in growing E. coli cells. We demonstrate that pre-existing lateral clusters tend to avoid translocation to pole regions and, therefore, continually shuttle between the cell poles for many generations while being static relative to the local cell-wall matrix. We also show that the polar preference of clusters results fundamentally from reduced clustering efficiency in the lateral region, rather than a developmental-like progression of clusters. Furthermore, polar preference is surprisingly robust to structural alterations designed to probe preference due to curvature sorting, perturbing the cell envelope physiology affects the cluster-size distribution, and the size-dependent mobility of receptor complexes differs between polar and lateral regions. Thus, distinct envelope physiology in the polar and lateral cell regions may contribute to polar preference.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

