3 minutes to precisely measure morphogen concentration
- PMID: 30365533
- PMCID: PMC6221364
- DOI: 10.1371/journal.pgen.1007676
3 minutes to precisely measure morphogen concentration
Abstract
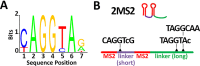
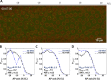
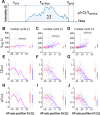
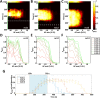
Morphogen gradients provide concentration-dependent positional information along polarity axes. Although the dynamics of the establishment of these gradients is well described, precision and noise in the downstream activation processes remain elusive. A simple paradigm to address these questions is the Bicoid morphogen gradient that elicits a rapid step-like transcriptional response in young fruit fly embryos. Focusing on the expression of the major Bicoid target, hunchback (hb), at the onset of zygotic transcription, we used the MS2-MCP approach which combines fluorescent labeling of nascent mRNA with live imaging at high spatial and temporal resolution. Removing 36 putative Zelda binding sites unexpectedly present in the original MS2 reporter, we show that the 750 bp of the hb promoter are sufficient to recapitulate endogenous expression at the onset of zygotic transcription. After each mitosis, in the anterior, expression is turned on to rapidly reach a plateau with all nuclei expressing the reporter. Consistent with a Bicoid dose-dependent activation process, the time period required to reach the plateau increases with the distance to the anterior pole. Despite the challenge imposed by frequent mitoses and high nuclei-to-nuclei variability in transcription kinetics, it only takes 3 minutes at each interphase for the MS2 reporter loci to distinguish subtle differences in Bicoid concentration and establish a steadily positioned and steep (Hill coefficient ~ 7) expression boundary. Modeling based on the cooperativity between the 6 known Bicoid binding sites in the hb promoter region, assuming rate limiting concentrations of the Bicoid transcription factor at the boundary, is able to capture the observed dynamics of pattern establishment but not the steepness of the boundary. This suggests that a simple model based only on the cooperative binding of Bicoid is not sufficient to describe the spatiotemporal dynamics of early hb expression.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



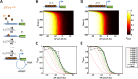
References
-
- Driever W, Nusslein-Volhard C. A gradient of bicoid protein in Drosophila embryos. Cell. 1988;54(1):83–93. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

