Mycoplasma genitalium adhesin P110 binds sialic-acid human receptors
- PMID: 30367053
- PMCID: PMC6203739
- DOI: 10.1038/s41467-018-06963-y
Mycoplasma genitalium adhesin P110 binds sialic-acid human receptors
Abstract
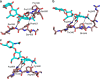
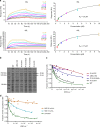
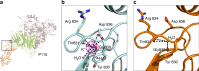
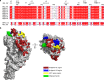
Adhesion of pathogenic bacteria to target cells is a prerequisite for colonization and further infection. The main adhesins of the emerging sexually transmitted pathogen Mycoplasma genitalium, P140 and P110, interact to form a Nap complex anchored to the cell membrane. Herein, we present the crystal structures of the extracellular region of the virulence factor P110 (916 residues) unliganded and in complex with sialic acid oligosaccharides. P110 interacts only with the neuraminic acid moiety of the oligosaccharides and experiments with human cells demonstrate that these interactions are essential for mycoplasma cytadherence. Additionally, structural information provides a deep insight of the P110 antigenic regions undergoing programmed variation to evade the host immune response. These results enlighten the interplay of M. genitalium with human target cells, offering new strategies to control mycoplasma infections.
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Dynamics of the adhesion complex of the human pathogens Mycoplasma pneumoniae and Mycoplasma genitalium.PLoS Pathog. 2025 Mar 28;21(3):e1012973. doi: 10.1371/journal.ppat.1012973. eCollection 2025 Mar. PLoS Pathog. 2025. PMID: 40153444 Free PMC article.
-
Stability of cytadherence-related proteins P140/P110 in Mycoplasma genitalium requires MG218 and unidentified factors.Arch Med Res. 2002 Jan-Feb;33(1):1-5. doi: 10.1016/s0188-4409(01)00335-6. Arch Med Res. 2002. PMID: 11825623
-
Mycoplasma genitalium P140 and P110 cytadhesins are reciprocally stabilized and required for cell adhesion and terminal-organelle development.J Bacteriol. 2006 Dec;188(24):8627-37. doi: 10.1128/JB.00978-06. Epub 2006 Oct 6. J Bacteriol. 2006. PMID: 17028283 Free PMC article.
-
Pathogenicity and virulence of Mycoplasma genitalium: Unraveling Ariadne's Thread.Virulence. 2022 Dec;13(1):1161-1183. doi: 10.1080/21505594.2022.2095741. Virulence. 2022. PMID: 35791283 Free PMC article. Review.
-
A structural overview of mycobacterial adhesins: Key biomarkers for diagnostics and therapeutics.Protein Sci. 2018 Feb;27(2):369-380. doi: 10.1002/pro.3346. Epub 2017 Dec 27. Protein Sci. 2018. PMID: 29139177 Free PMC article. Review.
Cited by
-
Sequence variation and immunogenicity of the Mycoplasma genitalium MgpB and MgpC adherence proteins during persistent infection of men with non-gonococcal urethritis.PLoS One. 2020 Oct 12;15(10):e0240626. doi: 10.1371/journal.pone.0240626. eCollection 2020. PLoS One. 2020. PMID: 33045031 Free PMC article.
-
Mbov_0503 Encodes a Novel Cytoadhesin that Facilitates Mycoplasma bovis Interaction with Tight Junctions.Microorganisms. 2020 Jan 23;8(2):164. doi: 10.3390/microorganisms8020164. Microorganisms. 2020. PMID: 31979335 Free PMC article.
-
Dynamics of the adhesion complex of the human pathogens Mycoplasma pneumoniae and Mycoplasma genitalium.PLoS Pathog. 2025 Mar 28;21(3):e1012973. doi: 10.1371/journal.ppat.1012973. eCollection 2025 Mar. PLoS Pathog. 2025. PMID: 40153444 Free PMC article.
-
Investigating the immunological activity of the Hsp70-P113 fusion protein for Mycoplasma ovipneumoniae detection: a groundbreaking study.BMC Vet Res. 2024 Sep 20;20(1):421. doi: 10.1186/s12917-024-04274-7. BMC Vet Res. 2024. PMID: 39304865 Free PMC article.
-
How bacteria utilize sialic acid during interactions with the host: snip, snatch, dispatch, match and attach.Microbiology (Reading). 2022 Mar;168(3):001157. doi: 10.1099/mic.0.001157. Microbiology (Reading). 2022. PMID: 35316172 Free PMC article. Review.

