Exploring the Role of AUG Triplets in Human Cap-Independent Translation Enhancing Elements
- PMID: 30371061
- PMCID: PMC6222554
- DOI: 10.1021/acs.biochem.8b00785
Exploring the Role of AUG Triplets in Human Cap-Independent Translation Enhancing Elements
Abstract
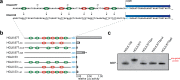
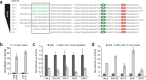
Cap-independent translation is believed to play an important role in eukaryotic protein synthesis, but the mechanisms of ribosomal recruitment and translation initiation remain largely unknown. Messenger RNA display was previously used to profile the human genome for RNA leader sequences that can enhance cap-independent translation. Surprisingly, many of the isolated sequences contain AUG triplets, suggesting a possible functional role for these motifs during translation initiation. Herein, we examine the sequence determinants of AUG triplets within a set of human translation enhancing elements (TEEs). Functional analyses performed in vitro and in cultured cells indicate that AUGs have the capacity to modulate mRNA translation either by serving as part of a larger ribosomal recruitment site or by directing the ribosome to defined initiation sites. These observations help constrain the functional role of AUG triplets in human TEEs and advance our understanding of this specific mechanism of cap-independent translation initiation.
Conflict of interest statement
The authors declare no competing financial interest.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

