Using human sequencing to guide craniofacial research
- PMID: 30375152
- PMCID: PMC8796141
- DOI: 10.1002/dvg.23259
Using human sequencing to guide craniofacial research
Abstract
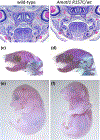
A recent convergence of technological innovations has re-energized the ability to apply genetics to research in human craniofacial development. Next-generation exome and whole genome sequencing have significantly dropped in price, making it relatively trivial to sequence and analyze patients and families with congenital craniofacial anomalies. A concurrent revolution in genome editing with the use of the CRISPR-Cas9 system enables the rapid generation of animal models, including mouse, which can precisely recapitulate human variants. Here, we summarize the choices currently available to the research community. We illustrate this approach with the study of a family with a novel craniofacial syndrome with dominant inheritance pattern. The genomic analysis suggested a causal variant in AMOTL1 which we modeled in mice. We also made a novel deletion allele of Amotl1. Our results indicate that Amotl1 is not required in the mouse for survival to weaning. Mice carrying the variant identified in the human sequencing studies, however, do not survive to weaning in normal ratios. The cause of death is not understood for these mice complicating our conclusions about the pathogenicity in the index patient. Thus, we highlight some of the powerful opportunities and confounding factors confronting current craniofacial genetic research.
Keywords: birth defects; mammal; organism; process, genetics; process, neural crest; process, organogenesis; tissue; tissue, other.
© 2018 Wiley Periodicals, Inc.
Figures

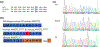


References
-
- Bratt A, Wilson WJ, Troyanovsky B, Aase K, Kessler R, Van Meir EG, & Holmgren L. (2002). Angiomotin belongs to a novel protein family with conserved coiled-coil and PDZ binding domains. Gene, 298, 69–77. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

