PIF4-induced BR synthesis is critical to diurnal and thermomorphogenic growth
- PMID: 30389669
- PMCID: PMC6276883
- DOI: 10.15252/embj.201899552
PIF4-induced BR synthesis is critical to diurnal and thermomorphogenic growth
Abstract
The Arabidopsis PIF4 and BES1/BZR1 transcription factors antagonize light signaling by facilitating co-activated expression of a large number of cell wall-loosening and auxin-related genes. While PIF4 directly activates expression of these targets, BES1 and BZR1 activity switch from a repressive to an activator function, depending on interaction with TOPLESS and other families of regulators including PIFs. However, the complexity of this regulation and its role in diurnal control of plant growth and brassinosteroid (BR) levels is little understood. We show by using a protein array that BES1, PIF4, and the BES1-PIF4 complex recognize different DNA elements, thus revealing a distinctive cis-regulatory code beneath BES1-repressive and PIF4 co-activation function. BES1 homodimers bind to conserved BRRE- and G-box elements in the BR biosynthetic promoters and inhibit their expression during the day, while elevated PIF4 competes for BES1 homodimer formation, resulting in de-repressed BR biosynthesis at dawn and in response to warmth. Our findings demonstrate a central role of PIF4 in BR synthesis activation, increased BR levels being essential to thermomorphogenic hypocotyl growth.
Keywords: Arabidopsis; BES1; BR biosynthesis; PIF4; thermomorphogenesis.
© 2018 The Authors.
Figures
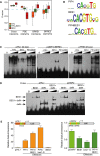
Box plot showing the enrichment scores (E‐scores) of all possible 8‐mers containing the G‐box (CACGTG), PBE‐box (CATGTG), and BRRE‐elements (CGTGC/TG). Boxes represent the 25–75% quartiles and the black line the median of distribution. Bars indicate the 1–25% (above) and 75–100% (below) quartiles. E‐scores above 0.4 denote that binding of the proteins toward the indicated DNA element is statistically significant. Dashed blue line indicates the 0.4 threshold.
Sequence logo representation of the top scoring 8‐mers obtained by hybridization with the PIF4, BES1, and PIF4‐BES1 proteins.
Electrophoretic mobility shift assays (EMSA) showing interaction of the PIF4, BES1, and PIF4‐BES1 proteins with the conserved G‐box, PBE‐, and BRRE‐elements in the PIL1 (At2g46970), PRE5 (At3g28857), and DWF4 (At3g50660) promoters. Increasing amounts of protein were used for the assay.
BES1 binds both BRRE‐ and G‐box elements as a homodimer. A deletion of BES1 (delN) fused to MBP (MBP‐delN) and the complete protein (MBP‐BES1) was co‐expressed in E. coli. Formation of intermediate mobility bands, indicative of DNA recognition by a dimeric form of the protein, was tested in EMSA assays. A signal corresponding to the dimeric full‐length MBP‐BES1 and MBP‐delN complex was detected with both DNA probes. Increasing amounts of the proteins were used in the assay.
Co‐expression of the PIF4 and BES1 effector constructs leads to synergistic pPIL1 activation and reverses BES1‐dependent inhibition of the pDWF4 reporter. The pPIL1 and pDWF4 promoters including three G‐boxes (green boxes) and two BRRE‐elements (orange boxes) were fused to the firefly luciferase reporter gene (LUC) and co‐transfected with 35S::PIF4, 35S::BES1, and 35S::bes1‐D effector constructs into Nicotiana benthamiana leaves. Leaf disks were collected 48 h after infiltration, and luciferase activity was measured in a microplate luminometer. Error bars represent SD (n = 24).
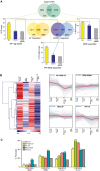
Venn diagrams representing the genes up‐regulated (yellow) and down‐regulated (purple) by PIFs or BES1/BZR1 (BES), and the direct target loci (green) of these transcription factors. Gene sets derive from our bes1‐D and bes1‐D;pif4pif5 microarray analyses and from published RNA‐seq studies of pifq, bzr1‐1D, and bri1‐116 mutants (Appendix Tables S1 and S2). The direct target PIF and BES datasets were obtained from published ChIP‐chip or ChIP‐seq experiments with these factors (Appendix Tables S3 and S4). Histogram bars represent the percentage of PIF‐, BES‐, and PIF + BES‐regulated genes that are directly bound by these transcription factors, as compared to all the Arabidopsis genes (total genes, in gray) (chi‐squared test, **P‐value < 0.01, ns indicates no significant differences).
Heat map and fold‐change graph representation of the genes identified in (A). Expression data were obtained from Genevestigator, and heat map and clustering analyses were performed using the MeV software. PIF + BES‐UP indicates genes up‐regulated by PIF and BES and identified as targets of both the PIF and BES factors. BES‐DOWN are BES‐repressed targets, while BES‐UP corresponds to BES‐induced targets. The PIF‐UP cluster includes PIF‐induced genes directly bound by the PIF factors.
Percentages of occurrence of the indicated motifs in the PIF, BES, or PIF + BES target loci, as compared with all Arabidopsis genes (genome, in gray) (chi‐squared test, **P‐value < 0.01, ns indicates no significant differences).
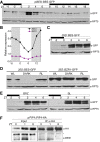
Time course analysis of BES1 protein accumulation levels. pBES1::BES1‐GFP seedlings were grown under short days for 6 days, and samples were harvested each 2 h for a 24‐h interval. Total protein extracts were used for Western blot analysis with an anti‐GFP antibody to detect the BES1‐GFP protein. Hybridization with anti‐RPT5 antibody is included as a loading control. Dark and light periods are represented as black and white bars, respectively. The experiment was repeated twice with similar results.
Diurnal oscillation of BES1 and BZR1 transcripts. Col‐0 seedlings were grown for 6 days under short days, and samples were harvested every 3 h. BES1 and BZR1 expression levels were determined by qRT–PCR analyses and normalized to PP2A. Relative levels are the mean of three technical replicates, and error bars represent standard deviation. The experiment was performed three times with similar results. Dark periods are indicated by shading.
Western blot analyses of 35S::BES‐GFP seedlings grown for 6 days under short days. Samples were collected every 2 h, starting 2 h before lights on (ZT‐2) to ZT6. Western blots were hybridized with an anti‐GFP antibody to detect the BES1‐GFP protein and anti‐RPT5 as a loading control.
BES1 but not BZR1 is stabilized in the light. 35S::BES1‐GFP and 35S::BZR1‐GFP seedlings were grown for 6 days in continuous white light (WL), red light (RL), and continuous dark (DARK), and the BES1 and BZR1 proteins analyzed by Western blot using an anti‐GFP antibody. Hybridization with an anti‐RPT5 antibody was used as a loading control.
BES1 accumulation is independent of brassinosteroid levels. Six‐day‐old 35S::BES‐GFP seedlings were treated overnight with 100 nM epi‐brassinolide (BL) or 0.5 μM brassinazole (BRZ). Material was collected at the indicated time points and analyzed by Western blot hybridization with anti‐GFP and anti‐RPT5 antibodies.
In vivo analysis of PIF4‐BES1 complex formation. Six‐day‐old pPIF4::PIF4‐HA seedlings grown under short‐day conditions were collected at ZT0, ZT1, and ZT8. PIF4‐HA was immunoprecipitated with an anti‐HA antibody and the pulled‐down BES1 protein immunodetected with an anti‐BES antibody. *indicates a non‐specific band by the primary antibody used to immunoprecipitation.
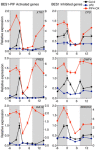
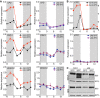
Col‐0 and pifq seedlings were grown under short days at 22 and 28°C, and samples were collected at the indicated time points. Expression of BR biosynthetic genes was analyzed by qRT–PCR, using the constitutive PP2A gene as internal control. Error bars represent SD of three technical replicates. The experiment was repeated twice with similar results.
Activation of the XTR7 PIF + BES‐UP target was studied as a positive control of the experiment. Error bars represent SD of three technical replicates. Dark periods are shown in gray. The experiment was performed three times with similar results.
Increased levels of dephosphorylated BES1 at elevated temperatures depend on PIFs activation of BR synthesis. Col‐0 and pifq seedlings were grown for 6 days under short days and 22°C or 28°C, on half strength MS media (mock) or media supplemented with 0.5 μM BRZ. BES1 was analyzed by Western blot hybridization with an anti‐BES1 antibody. RPT5 was used as a loading control.
Similar articles
-
BR-dependent phosphorylation modulates PIF4 transcriptional activity and shapes diurnal hypocotyl growth.Genes Dev. 2014 Aug 1;28(15):1681-94. doi: 10.1101/gad.243675.114. Genes Dev. 2014. PMID: 25085420 Free PMC article.
-
Brassinosteroids Dominate Hormonal Regulation of Plant Thermomorphogenesis via BZR1.Curr Biol. 2018 Jan 22;28(2):303-310.e3. doi: 10.1016/j.cub.2017.11.077. Epub 2018 Jan 11. Curr Biol. 2018. PMID: 29337075
-
Transcription factor HAT1 is phosphorylated by BIN2 kinase and mediates brassinosteroid repressed gene expression in Arabidopsis.Plant J. 2014 Jan;77(1):59-70. doi: 10.1111/tpj.12368. Epub 2013 Dec 3. Plant J. 2014. PMID: 24164091
-
The brassinosteroid-regulated transcription factors BZR1/BES1 function as a coordinator in multisignal-regulated plant growth.Biochim Biophys Acta Gene Regul Mech. 2018 Jun;1861(6):561-571. doi: 10.1016/j.bbagrm.2018.04.003. Epub 2018 Apr 17. Biochim Biophys Acta Gene Regul Mech. 2018. PMID: 29673687 Review.
-
Interaction of the Transcription Factors BES1/BZR1 in Plant Growth and Stress Response.Int J Mol Sci. 2024 Jun 21;25(13):6836. doi: 10.3390/ijms25136836. Int J Mol Sci. 2024. PMID: 38999944 Free PMC article. Review.
Cited by
-
How plants coordinate their development in response to light and temperature signals.Plant Cell. 2022 Mar 4;34(3):955-966. doi: 10.1093/plcell/koab302. Plant Cell. 2022. PMID: 34904672 Free PMC article. Review.
-
Warm temperature suppresses plant systemic acquired resistance by intercepting N-hydroxypipecolic acid biosynthesis.Plant J. 2025 Aug;123(3):e70374. doi: 10.1111/tpj.70374. Plant J. 2025. PMID: 40758777 Free PMC article.
-
Abiotic Stress Tolerance in Plants: Brassinosteroids Navigate Competently.Int J Mol Sci. 2022 Nov 23;23(23):14577. doi: 10.3390/ijms232314577. Int J Mol Sci. 2022. PMID: 36498906 Free PMC article. Review.
-
Inhibitors of Brassinosteroid Biosynthesis and Signal Transduction.Molecules. 2019 Nov 29;24(23):4372. doi: 10.3390/molecules24234372. Molecules. 2019. PMID: 31795392 Free PMC article. Review.
-
Exogenous application of β-sitosterol mediated growth and yield improvement in water-stressed wheat (Triticum aestivum) involves up-regulated antioxidant system.J Plant Res. 2019 Nov;132(6):881-901. doi: 10.1007/s10265-019-01143-5. Epub 2019 Oct 12. J Plant Res. 2019. PMID: 31606785
References
-
- Box MS, Huang BE, Domijan M, Jaeger KE, Khattak AK, Yoo SJ, Sedivy EL, Jones DM, Hearn TJ, Webb AA, Grant A, Locke JC, Wigge PA (2015) ELF3 controls thermoresponsive growth in Arabidopsis . Curr Biol 25: 194–199 - PubMed
-
- Espinosa‐Ruiz A, Martinez C, de Lucas M, Fabregas N, Bosch N, Cano‐Delgado AI, Prat S (2017) TOPLESS mediates brassinosteroid control of shoot boundaries and root meristem development in Arabidopsis thaliana . Development 144: 1619–1628 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

