Nanowell-mediated multidimensional separations combining nanoLC with SLIM IM-MS for rapid, high-peak-capacity proteomic analyses
- PMID: 30397757
- PMCID: PMC6500776
- DOI: 10.1007/s00216-018-1452-5
Nanowell-mediated multidimensional separations combining nanoLC with SLIM IM-MS for rapid, high-peak-capacity proteomic analyses
Abstract
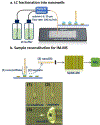
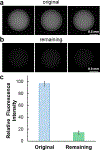
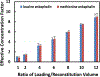
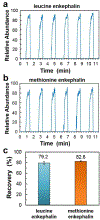
Mass spectrometry (MS)-based analysis of complex biological samples is essential for biomedical research and clinical diagnostics. The separation prior to MS plays a key role in the overall analysis, with separations having larger peak capacities often leading to more identified species and improved confidence in those identifications. High-resolution ion mobility (IM) separations enabled by Structures for Lossless Ion Manipulation (SLIM) can provide extremely rapid, high-resolution separations and are well suited as a second dimension of separation following nanoscale liquid chromatography (nanoLC). However, existing sample handling approaches for offline coupling of separation modes require microliter-fraction volumes and are thus not well suited for analysis of trace biological samples. We have developed a novel nanowell-mediated fractionation system that enables nanoLC-separated samples to be efficiently preconcentrated and directly infused at nanoelectrospray flow rates for downstream analysis. When coupled with SLIM IM-MS, the platform enables rapid and high-peak-capacity multidimensional separations of small biological samples. In this study, peptides eluting from a 100 nL/min nanoLC separation were fractionated into ~ 60 nanowells on a microfluidic glass chip using an in-house-developed robotic system. The dried samples on the chip were individually reconstituted and ionized by nanoelectrospray for SLIM IM-MS analysis. Using model peptides for characterization of the nanowell platform, we found that at least 80% of the peptide components of the fractionated samples were recovered from the nanowells, providing up to ~tenfold preconcentration for SLIM IM-MS analysis. The combined LC-SLIM IM separation peak capacities exceeded 3600 with a measurement throughput that is similar to current one-dimensional (1D) LC-MS proteomic analyses. Graphical abstract A nanowell-mediated multidimensional separation platform that combines nanoLC with SLIM IM-MS enables rapid, high-peak-capacity proteomic analyses.
Keywords: Ion mobility; Mass spectrometry; Nanoelectrospray; nanoPOTS.
Conflict of interest statement
The authors declare no conflict of interests.
Figures
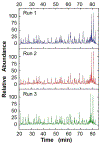
References
-
- Ong S-E, Mann M (2005) Mass spectrometry–based proteomics turns quantitative. Nat. Chem. Biol 1 (5):252–262 - PubMed
-
- Bantscheff M, Schirle M, Sweetman G, Rick J, Kuster B (2007) Quantitative mass spectrometry in proteomics: a critical review. Anal. Bioanal. Chem 389 (4):1017–1031 - PubMed
-
- Boschetti E, Righetti PG (2013) Low-abundance proteome discovery: state of the art and protocols. Elsevier, Oxford UK.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

