Modeling Skeletal Muscle Laminopathies Using Human Induced Pluripotent Stem Cells Carrying Pathogenic LMNA Mutations
- PMID: 30405424
- PMCID: PMC6201196
- DOI: 10.3389/fphys.2018.01332
Modeling Skeletal Muscle Laminopathies Using Human Induced Pluripotent Stem Cells Carrying Pathogenic LMNA Mutations
Abstract
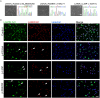
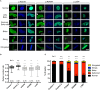
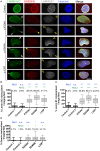
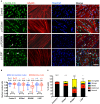
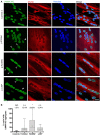
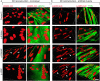
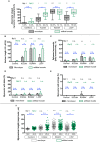
Laminopathies are a clinically heterogeneous group of disorders caused by mutations in LMNA. The main proteins encoded by LMNA are Lamin A and C, which together with Lamin B1 and B2, form the nuclear lamina: a mesh-like structure located underneath the inner nuclear membrane. Laminopathies show striking tissue specificity, with subtypes affecting striated muscle, peripheral nerve, and adipose tissue, while others cause multisystem disease with accelerated aging. Although several pathogenic mechanisms have been proposed, the exact pathophysiology of laminopathies remains unclear, compounded by the rarity of these disorders and lack of easily accessible cell types to study. To overcome this limitation, we used induced pluripotent stem cells (iPSCs) from patients with skeletal muscle laminopathies such as LMNA-related congenital muscular dystrophy and limb-girdle muscular dystrophy 1B, to model disease phenotypes in vitro. iPSCs can be derived from readily accessible cell types, have unlimited proliferation potential and can be differentiated into cell types that would otherwise be difficult and invasive to obtain. iPSC lines from three skeletal muscle laminopathy patients were differentiated into inducible myogenic cells and myotubes. Disease-associated phenotypes were observed in these cells, including abnormal nuclear shape and mislocalization of nuclear lamina proteins. Nuclear abnormalities were less pronounced in monolayer cultures of terminally differentiated skeletal myotubes than in proliferating myogenic cells. Notably, skeletal myogenic differentiation of LMNA-mutant iPSCs in artificial muscle constructs improved detection of myonuclear abnormalities compared to conventional monolayer cultures across multiple pathogenic genotypes, providing a high-fidelity modeling platform for skeletal muscle laminopathies. Our results lay the foundation for future iPSC-based therapy development and screening platforms for skeletal muscle laminopathies.
Keywords: 3D modeling; LMNA; disease modeling; iPSCs; lamin A/C; laminopathies; muscular dystrophy; skeletal muscle.
Figures
Similar articles
-
Lamin-Related Congenital Muscular Dystrophy Alters Mechanical Signaling and Skeletal Muscle Growth.Int J Mol Sci. 2020 Dec 30;22(1):306. doi: 10.3390/ijms22010306. Int J Mol Sci. 2020. PMID: 33396724 Free PMC article.
-
A CRISPR/Cas9 zebrafish lamin A/C mutant model of muscular laminopathy.Dev Dyn. 2022 Apr;251(4):645-661. doi: 10.1002/dvdy.427. Epub 2021 Oct 18. Dev Dyn. 2022. PMID: 34599606
-
Characteristics of nuclear architectural abnormalities of myotubes differentiated from LmnaH222P/H222P skeletal muscle cells.In Vitro Cell Dev Biol Anim. 2024 Aug;60(7):781-792. doi: 10.1007/s11626-024-00915-1. Epub 2024 May 9. In Vitro Cell Dev Biol Anim. 2024. PMID: 38724872
-
Induced Pluripotent Stem Cells to Study Mechanisms of Laminopathies: Focus on Epigenetics.Front Cell Dev Biol. 2018 Dec 20;6:172. doi: 10.3389/fcell.2018.00172. eCollection 2018. Front Cell Dev Biol. 2018. PMID: 30619852 Free PMC article. Review.
-
Need for NAD+: Focus on Striated Muscle Laminopathies.Cells. 2020 Oct 7;9(10):2248. doi: 10.3390/cells9102248. Cells. 2020. PMID: 33036437 Free PMC article. Review.
Cited by
-
Challenging the "chromatin hypothesis" of cardiac laminopathies with LMNA mutant iPS cells.J Cell Biol. 2019 Sep 2;218(9):2826-2828. doi: 10.1083/jcb.201907166. Epub 2019 Aug 19. J Cell Biol. 2019. PMID: 31427369 Free PMC article.
-
Advanced models of human skeletal muscle differentiation, development and disease: Three-dimensional cultures, organoids and beyond.Curr Opin Cell Biol. 2021 Dec;73:92-104. doi: 10.1016/j.ceb.2021.06.004. Epub 2021 Aug 9. Curr Opin Cell Biol. 2021. PMID: 34384976 Free PMC article. Review.
-
Versatility of Induced Pluripotent Stem Cells (iPSCs) for Improving the Knowledge on Musculoskeletal Diseases.Int J Mol Sci. 2020 Aug 25;21(17):6124. doi: 10.3390/ijms21176124. Int J Mol Sci. 2020. PMID: 32854405 Free PMC article. Review.
-
Induced Pluripotent Stem Cells for Modeling Physiological and Pathological Striated Muscle Complexity.J Neuromuscul Dis. 2023;10(5):761-776. doi: 10.3233/JND-230076. J Neuromuscul Dis. 2023. PMID: 37522215 Free PMC article. Review.
-
Advanced Cellular Models for Rare Disease Study: Exploring Neural, Muscle and Skeletal Organoids.Int J Mol Sci. 2024 Jan 13;25(2):1014. doi: 10.3390/ijms25021014. Int J Mol Sci. 2024. PMID: 38256087 Free PMC article. Review.
References
-
- Arimura T., Helbling-Leclerc A., Massart C., Varnous S., Niel F., Lacène E., et al. . (2005). Mouse model carrying H222P-Lmna mutation develops muscular dystrophy and dilated cardiomyopathy similar to human striated muscle laminopathies. Hum. Mol. Genet. 14, 155–169. 10.1093/hmg/ddi017 - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

