Functional heterogeneity of human tissue-resident memory T cells based on dye efflux capacities
- PMID: 30429372
- PMCID: PMC6302950
- DOI: 10.1172/jci.insight.123568
Functional heterogeneity of human tissue-resident memory T cells based on dye efflux capacities
Abstract
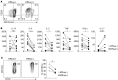
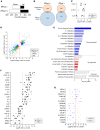
Tissue-resident memory T cells (TRMs) accelerate pathogen clearance through rapid and enhanced functional responses in situ. TRMs are prevalent in diverse anatomic sites throughout the human lifespan, yet their phenotypic and functional diversity has not been fully described. Here, we identify subpopulations of human TRMs based on the ability to efflux fluorescent dyes [efflux(+) TRMs] located within mucosal and lymphoid sites with distinct transcriptional profiles, turnover, and functional capacities. Compared with efflux(-) TRMs, efflux(+) TRMs showed transcriptional and phenotypic features of quiescence including reduced turnover, decreased expression of exhaustion markers, and increased proliferative capacity and signaling in response to homeostatic cytokines. Moreover, upon activation, efflux(+) TRMs secreted lower levels of inflammatory cytokines such as IFN-γ and IL-2 and underwent reduced degranulation. Interestingly, analysis of TRM subsets following activation revealed that both efflux(+) and efflux(-) TRMs undergo extensive transcriptional changes following TCR ligation but retain core TRM transcriptional properties including retention markers, suggesting that TRMs carry out effector function in situ. Overall, our results suggest a model for tissue-resident immunity wherein heterogeneous subsets have differential capacities for longevity and effector function.
Keywords: Adaptive immunity; Immunology; T cells.
Conflict of interest statement
Figures





References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

