Gut and blood differ in constitutive blocks to HIV transcription, suggesting tissue-specific differences in the mechanisms that govern HIV latency
- PMID: 30440043
- PMCID: PMC6237391
- DOI: 10.1371/journal.ppat.1007357
Gut and blood differ in constitutive blocks to HIV transcription, suggesting tissue-specific differences in the mechanisms that govern HIV latency
Abstract
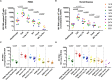
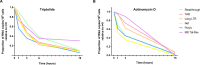
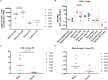
Latently-infected CD4+ T cells are widely considered to be the major barrier to a cure for HIV. Much of our understanding of HIV latency comes from latency models and blood cells, but most HIV-infected cells reside in lymphoid tissues such as the gut. We hypothesized that tissue-specific environments may impact the mechanisms that govern HIV expression. To assess the degree to which different mechanisms inhibit HIV transcription in the gut and blood, we quantified HIV transcripts suggestive of transcriptional interference (U3-U5; "Read-through"), initiation (TAR), 5' elongation (R-U5-pre-Gag; "Long LTR"), distal transcription (Nef), completion (U3-polyA; "PolyA"), and multiple splicing (Tat-Rev) in matched peripheral blood mononuclear cells (PBMCs) and rectal biopsies, and matched FACS-sorted CD4+ T cells from blood and rectum, from two cohorts of ART-suppressed individuals. Like the PBMCs, rectal biopsies showed low levels of read-through transcripts (median = 23 copies/106 cells) and a gradient of total (679)>elongated(75)>Nef(16)>polyadenylated (11)>multiply-spliced HIV RNAs(<1) [p<0.05 for all], demonstrating blocks to HIV transcriptional elongation, completion, and splicing. Rectal CD4+ T cells showed a similar gradient of total>polyadenylated>multiply-spliced transcripts, but the ratio of total to elongated transcripts was 6-fold lower than in blood CD4+ T cells (P = 0.016), suggesting less of a block to HIV transcriptional elongation in rectal CD4+ T cells. Levels of total transcripts per provirus were significantly lower in rectal biopsies compared to PBMCs (median 3.5 vs. 15.4; P = 0.008) and in sorted CD4+ T cells from rectum compared to blood (median 2.7 vs. 31.8; P = 0.016). The lower levels of HIV transcriptional initiation and of most HIV transcripts per provirus in the rectum suggest that this site may be enriched for latently-infected cells, cells in which latency is maintained by different mechanisms, or cells in a "deeper" state of latency. These are important considerations for designing therapies that aim to disrupt HIV latency in all tissue compartments.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


References
-
- Chun TW, Carruth L, Finzi D, Shen X, DiGiuseppe JA, Taylor H, et al. Quantification of latent tissue reservoirs and total body viral load in HIV-1 infection. Nature. 1997;387(6629):183–8. 10.1038/387183a0 . - DOI - PubMed
-
- Finzi D, Hermankova M, Pierson T, Carruth LM, Buck C, Chaisson RE, et al. Identification of a reservoir for HIV-1 in patients on highly active antiretroviral therapy. Science. 1997;278(5341):1295–300. . - PubMed
-
- Wong JK, Hezareh M, Gunthard HF, Havlir DV, Ignacio CC, Spina CA, et al. Recovery of replication-competent HIV despite prolonged suppression of plasma viremia. Science. 1997;278(5341):1291–5. . - PubMed
-
- Chomont N, El-Far M, Ancuta P, Trautmann L, Procopio FA, Yassine-Diab B, et al. HIV reservoir size and persistence are driven by T cell survival and homeostatic proliferation. Nat Med. 2009;15(8):893–900. 10.1038/nm.1972 ; PubMed Central PMCID: PMCPMC2859814. - DOI - PMC - PubMed
-
- Finzi D, Blankson J, Siliciano JD, Margolick JB, Chadwick K, Pierson T, et al. Latent infection of CD4+ T cells provides a mechanism for lifelong persistence of HIV-1, even in patients on effective combination therapy. Nature Medicine. 1999;5(5):512–7. Epub 1999/05/06. 10.1038/8394 . - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

