An aromatic amino acid and associated helix in the C-terminus of the potato leafroll virus minor capsid protein regulate systemic infection and symptom expression
- PMID: 30440046
- PMCID: PMC6264904
- DOI: 10.1371/journal.ppat.1007451
An aromatic amino acid and associated helix in the C-terminus of the potato leafroll virus minor capsid protein regulate systemic infection and symptom expression
Abstract
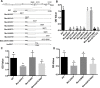
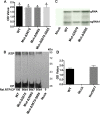
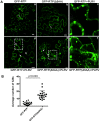
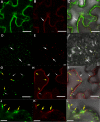
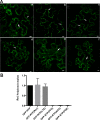
The C-terminal region of the minor structural protein of potato leafroll virus (PLRV), known as the readthrough protein (RTP), is involved in efficient virus movement, tissue tropism and symptom development. Analysis of numerous C-terminal deletions identified a five-amino acid motif that is required for RTP function. A PLRV mutant expressing RTP with these five amino acids deleted (Δ5aa-RTP) was compromised in systemic infection and symptom expression. Although the Δ5aa-RTP mutant was able to move long distance, limited infection foci were observed in systemically infected leaves suggesting that these five amino acids regulate virus phloem loading in the inoculated leaves and/or unloading into the systemically infected tissues. The 5aa deletion did not alter the efficiency of RTP translation, nor impair RTP self-interaction or its interaction with P17, the virus movement protein. However, the deletion did alter the subcellular localization of RTP. When co-expressed with a PLRV infectious clone, a GFP tagged wild-type RTP was localized to discontinuous punctate spots along the cell periphery and was associated with plasmodesmata, although localization was dependent upon the developmental stage of the plant tissue. In contrast, the Δ5aa-RTP-GFP aggregated in the cytoplasm. Structural modeling indicated that the 5aa deletion would be expected to perturb an α-helix motif. Two of 30 plants infected with Δ5aa-RTP developed a wild-type virus infection phenotype ten weeks post-inoculation. Analysis of the virus population in these plants by deep sequencing identified a duplication of sequences adjacent to the deletion that were predicted to restore the α-helix motif. The subcellular distribution of the RTP is regulated by the 5-aa motif which is under strong selection pressure and in turn contributes to the efficient long distance movement of the virus and the induction of systemic symptoms.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Similar articles
-
A Stem-Loop Structure in Potato Leafroll Virus Open Reading Frame 5 (ORF5) Is Essential for Readthrough Translation of the Coat Protein ORF Stop Codon 700 Bases Upstream.J Virol. 2018 May 14;92(11):e01544-17. doi: 10.1128/JVI.01544-17. Print 2018 Jun 1. J Virol. 2018. PMID: 29514911 Free PMC article.
-
Small deletions in the potato leafroll virus readthrough protein affect particle morphology, aphid transmission, virus movement and accumulation.J Gen Virol. 2008 Aug;89(Pt 8):2037-2045. doi: 10.1099/vir.0.83625-0. J Gen Virol. 2008. PMID: 18632976
-
Host-dependent requirement for the Potato leafroll virus 17-kda protein in virus movement.Mol Plant Microbe Interact. 2002 Oct;15(10):1086-94. doi: 10.1094/MPMI.2002.15.10.1086. Mol Plant Microbe Interact. 2002. PMID: 12437307
-
Cross-linking measurements of the Potato leafroll virus reveal protein interaction topologies required for virion stability, aphid transmission, and virus-plant interactions.J Proteome Res. 2012 May 4;11(5):2968-81. doi: 10.1021/pr300041t. Epub 2012 Mar 30. J Proteome Res. 2012. PMID: 22390342 Free PMC article.
-
Potato leafroll virus structural proteins manipulate overlapping, yet distinct protein interaction networks during infection.Proteomics. 2015 Jun;15(12):2098-112. doi: 10.1002/pmic.201400594. Epub 2015 Apr 29. Proteomics. 2015. PMID: 25787689
Cited by
-
Synergism Among the Four Tobacco Bushy Top Disease Casual Agents in Symptom Induction and Aphid Transmission.Front Microbiol. 2022 Apr 4;13:846857. doi: 10.3389/fmicb.2022.846857. eCollection 2022. Front Microbiol. 2022. PMID: 35444628 Free PMC article.
-
Five questions on the cell-to-cell movement of Orthotospoviruses.BBA Adv. 2024 Oct 16;6:100124. doi: 10.1016/j.bbadva.2024.100124. eCollection 2024. BBA Adv. 2024. PMID: 39498475 Free PMC article.
-
The Carboxyl Terminal Regions of P0 Protein Are Required for Systemic Infections of Poleroviruses.Int J Mol Sci. 2022 Feb 9;23(4):1945. doi: 10.3390/ijms23041945. Int J Mol Sci. 2022. PMID: 35216065 Free PMC article.
-
Complex and simple translational readthrough signals in pea enation mosaic virus 1 and potato leafroll virus, respectively.PLoS Pathog. 2022 Sep 29;18(9):e1010888. doi: 10.1371/journal.ppat.1010888. eCollection 2022 Sep. PLoS Pathog. 2022. PMID: 36174104 Free PMC article.
-
Discovery of aphid-transmitted Rice tiller inhibition virus from native plants through metagenomic sequencing.PLoS Pathog. 2023 Mar 24;19(3):e1011238. doi: 10.1371/journal.ppat.1011238. eCollection 2023 Mar. PLoS Pathog. 2023. PMID: 36961862 Free PMC article.
References
-
- Gray S, Gildow FE. Luteovirus-aphid interactions. Annu Rev Phytopathol. 2003;41:539–66. Epub 2003/05/06. 10.1146/annurev.phyto.41.012203.105815 . - DOI - PubMed
-
- Gray S, Cilia M, Ghanim M. Circulative, "nonpropagative" virus transmission: an orchestra of virus-, insect-, and plant-derived instruments. Adv Virus Res. 2014;89:141–99. Epub 2014/04/23. 10.1016/B978-0-12-800172-1.00004-5 . - DOI - PubMed
-
- Hipper C, Monsion B, Bortolamiol-Becet D, Ziegler-Graff V, Brault V. Formation of virions is strictly required for Turnip yellows virus long-distance movement in plants. J Gen Virol. 2014;95:496–505. 10.1099/vir.0.058867-0 WOS:000338177400027. - DOI - PubMed
-
- Lee L, Kaplan IB, Ripoll DR, Liang D, Palukaitis P, Gray SM. A surface loop of the Potato leafroll virus coat protein is involved in virion assembly, systemic movement, and aphid transmission. J Virol. 2005;79(2):1207–14. 10.1128/JVI.79.2.1207-1214.2005 . - DOI - PMC - PubMed
-
- Schmitz J, Stussi-Garaud C, Tacke E, Prufer D, Rohde W, Rohfritsch O. In situ localization of the putative movement protein (pr17) from Potato leafroll luteovirus (PLRV) in infected and transgenic potato plants. Virology. 1997;235(2):311–22. 10.1006/viro.1997.8679 . - DOI - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources

