Mutations in LZTR1 drive human disease by dysregulating RAS ubiquitination
- PMID: 30442762
- PMCID: PMC8058620
- DOI: 10.1126/science.aap7607
Mutations in LZTR1 drive human disease by dysregulating RAS ubiquitination
Abstract
The leucine zipper-like transcriptional regulator 1 (LZTR1) protein, an adaptor for cullin 3 (CUL3) ubiquitin ligase complex, is implicated in human disease, yet its mechanism of action remains unknown. We found that Lztr1 haploinsufficiency in mice recapitulates Noonan syndrome phenotypes, whereas LZTR1 loss in Schwann cells drives dedifferentiation and proliferation. By trapping LZTR1 complexes from intact mammalian cells, we identified the guanosine triphosphatase RAS as a substrate for the LZTR1-CUL3 complex. Ubiquitome analysis showed that loss of Lztr1 abrogated Ras ubiquitination at lysine-170. LZTR1-mediated ubiquitination inhibited RAS signaling by attenuating its association with the membrane. Disease-associated LZTR1 mutations disrupted either LZTR1-CUL3 complex formation or its interaction with RAS proteins. RAS regulation by LZTR1-mediated ubiquitination provides an explanation for the role of LZTR1 in human disease.
Copyright © 2018 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Conflict of interest statement
Figures
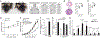


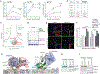
Comment in
-
Ras tuning.Nat Chem Biol. 2019 Feb;15(2):95. doi: 10.1038/s41589-018-0219-9. Nat Chem Biol. 2019. PMID: 30659296 No abstract available.
References
-
- Cancer Genome Atlas Research Network, Cell 169, 1327–1341. e23 (2017). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

