Orthogonal Cas9-Cas9 chimeras provide a versatile platform for genome editing
- PMID: 30451839
- PMCID: PMC6242970
- DOI: 10.1038/s41467-018-07310-x
Orthogonal Cas9-Cas9 chimeras provide a versatile platform for genome editing
Erratum in
-
Publisher Correction: Orthogonal Cas9-Cas9 chimeras provide a versatile platform for genome editing.Nat Commun. 2018 Dec 10;9(1):5294. doi: 10.1038/s41467-018-07776-9. Nat Commun. 2018. PMID: 30531933 Free PMC article.
Abstract
The development of robust, versatile and accurate toolsets is critical to facilitate therapeutic genome editing applications. Here we establish RNA-programmable Cas9-Cas9 chimeras, in single- and dual-nuclease formats, as versatile genome engineering systems. In both of these formats, Cas9-Cas9 fusions display an expanded targeting repertoire and achieve highly specific genome editing. Dual-nuclease Cas9-Cas9 chimeras have distinct advantages over monomeric Cas9s including higher target site activity and the generation of predictable precise deletion products between their target sites. At a therapeutically relevant site within the BCL11A erythroid enhancer, Cas9-Cas9 nucleases produced precise deletions that comprised up to 97% of all sequence alterations. Thus Cas9-Cas9 chimeras represent an important tool that could be particularly valuable for therapeutic genome editing applications where a precise cleavage position and defined sequence end products are desirable.
Conflict of interest statement
The authors declare the following competing interests: The authors have filed patent applications related to genome engineering technologies; E.J.S. is a co-founder and advisor of Intellia Therapeutics; S.A.W. is a consultant for Beam Therapeutics.
Figures

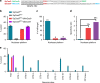




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

