Zebrafish Granulocyte Colony-Stimulating Factor Receptor Maintains Neutrophil Number and Function throughout the Life Span
- PMID: 30455199
- PMCID: PMC6346135
- DOI: 10.1128/IAI.00793-18
Zebrafish Granulocyte Colony-Stimulating Factor Receptor Maintains Neutrophil Number and Function throughout the Life Span
Abstract
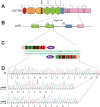
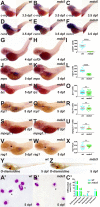
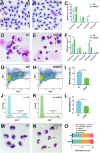
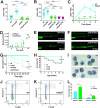
Granulocyte colony-stimulating factor receptor (G-CSFR), encoded by the CSF3R gene, represents a major regulator of neutrophil production and function in mammals, with inactivating extracellular mutations identified in a cohort of neutropenia patients unresponsive to G-CSF treatment. This study sought to elucidate the role of the zebrafish G-CSFR by generating mutants harboring these inactivating extracellular mutations using genome editing. Zebrafish csf3r mutants possessed significantly decreased numbers of neutrophils from embryonic to adult stages, which were also functionally compromised, did not respond to G-CSF, and displayed enhanced susceptibility to bacterial infection. The study has identified an important role for the zebrafish G-CSFR in maintaining the number and functionality of neutrophils throughout the life span and created a bona fide zebrafish model of nonresponsive neutropenia.
Keywords: blood diseases; cytokine receptors; developmental hematopoiesis; neutropenia; zebrafish.
Copyright © 2019 American Society for Microbiology.
Figures

Similar articles
-
Leukemia-associated truncation of granulocyte colony-stimulating factor receptor impacts granulopoiesis throughout the life-course.Front Immunol. 2023 Jan 10;13:1095453. doi: 10.3389/fimmu.2022.1095453. eCollection 2022. Front Immunol. 2023. PMID: 36703974 Free PMC article.
-
A GCSFR/CSF3R zebrafish mutant models the persistent basal neutrophil deficiency of severe congenital neutropenia.Sci Rep. 2017 Mar 10;7:44455. doi: 10.1038/srep44455. Sci Rep. 2017. PMID: 28281657 Free PMC article.
-
Increased granulocyte colony-stimulating factor responsiveness but normal resting granulopoiesis in mice carrying a targeted granulocyte colony-stimulating factor receptor mutation derived from a patient with severe congenital neutropenia.J Clin Invest. 1998 Aug 1;102(3):483-92. doi: 10.1172/JCI3216. J Clin Invest. 1998. PMID: 9691084 Free PMC article.
-
Granulocyte-macrophage colony-stimulating factor (GM-CSF) and granulocyte colony-stimulating factor (G-CSF): receptor biology, signal transduction, and neutrophil activation.Blood Rev. 1992 Mar;6(1):43-57. doi: 10.1016/0268-960x(92)90007-d. Blood Rev. 1992. PMID: 1375123 Review.
-
G-CSF, the guardian of granulopoiesis.Semin Immunol. 2021 Apr;54:101515. doi: 10.1016/j.smim.2021.101515. Epub 2021 Nov 10. Semin Immunol. 2021. PMID: 34772606 Review.
Cited by
-
Drainage of inflammatory macromolecules from the brain to periphery targets the liver for macrophage infiltration.Elife. 2020 Jul 31;9:e58191. doi: 10.7554/eLife.58191. Elife. 2020. PMID: 32735214 Free PMC article.
-
A zebrafish model for HAX1-associated congenital neutropenia.Haematologica. 2021 May 1;106(5):1311-1320. doi: 10.3324/haematol.2019.240200. Haematologica. 2021. PMID: 32327498 Free PMC article.
-
Stat3 Regulates Developmental Hematopoiesis and Impacts Myeloid Cell Function via Canonical and Non-Canonical Modalities.J Innate Immun. 2024;16(1):262-282. doi: 10.1159/000538364. Epub 2024 Apr 24. J Innate Immun. 2024. PMID: 38643762 Free PMC article.
-
Socs3b regulates the development and function of innate immune cells in zebrafish.Front Immunol. 2023 Mar 8;14:1119727. doi: 10.3389/fimmu.2023.1119727. eCollection 2023. Front Immunol. 2023. PMID: 36969252 Free PMC article.
-
A JAGN1-associated severe congenital neutropenia zebrafish model revealed an altered G-CSFR signaling and UPR activation.Blood Adv. 2024 Aug 13;8(15):4050-4065. doi: 10.1182/bloodadvances.2023011656. Blood Adv. 2024. PMID: 38739706 Free PMC article.
References
-
- Dale DC, Bolyard AA, Schwinzer BG, Pracht G, Bonilla MA, Boxer LA, Freedman MH, Donadieu J, Kannourakis G, Alter BP, Cham BP, Winkelstein J, Kinsey SE, Zeidler C, Welte K. 2006. The severe congenital neutropenia international registry: 10-year follow-up report. Support Cancer Ther 3:220–231. doi:10.3816/SCT.2006.n.020. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

