Overview of the Germline and Expressed Repertoires of the TRB Genes in Sus scrofa
- PMID: 30455691
- PMCID: PMC6230588
- DOI: 10.3389/fimmu.2018.02526
Overview of the Germline and Expressed Repertoires of the TRB Genes in Sus scrofa
Abstract
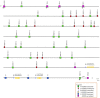
The α/β T cell receptor (TR) is a complex heterodimer that recognizes antigenic peptides and binds to major histocompatibility complex (MH) molecules. Both α and β chains are encoded by different genes localized on two distinct chromosomal loci: TRA and TRB. The present study employed the recent release of the swine genome assembly to define the genomic organization of the TRB locus. According to the sequencing data, the pig TRB locus spans approximately 400 kb of genomic DNA and consists of 38 TRBV genes belonging to 24 subgroups located upstream of three in tandem TRBD-J-C clusters, which are followed by a TRBV gene in an inverted transcriptional orientation. Comparative analysis confirms that the general organization of the TRB locus is similar among mammalian species, but the number of germline TRBV genes varies greatly even between species belonging to the same order, determining the diversity and specificity of the immune response. However, sequence analysis of the TRB locus also suggests the presence of blocks of conserved homology in the genomic region across mammals. Furthermore, by analysing a public cDNA collection, we identified the usage pattern of the TRBV, TRBD, and TRBJ genes in the adult pig TRB repertoire, and we noted that the expressed TRBV repertoire seems to be broader and more diverse than the germline repertoire, in line with the presence of a high level of TRBV gene polymorphisms. Because the nucleotide differences seems to be principally concentrated in the CDR2 region, it is reasonable to presume that most T cell β-chain diversity can be related to polymorphisms in pig MH molecules. Domestic pigs represent a valuable animal model as they are even more anatomically, genetically and physiologically similar to humans than are mice. Therefore, present knowledge on the genomic organization of the pig TRB locus allows the collection of increased information on the basic aspects of the porcine immune system and contributes to filling the gaps left by rodent models.
Keywords: IMGT; T cell receptor; TRB locus; TRB repertoire; domestic pig.
Figures






Similar articles
-
A Comprehensive Annotation of the Channel Catfish (Ictalurus punctatus) T Cell Receptor Alpha/Delta, Beta, and Gamma Loci.Front Immunol. 2021 Nov 25;12:786402. doi: 10.3389/fimmu.2021.786402. eCollection 2021. Front Immunol. 2021. PMID: 34899754 Free PMC article.
-
Genomic analysis reveals extensive gene duplication within the bovine TRB locus.BMC Genomics. 2009 Apr 24;10:192. doi: 10.1186/1471-2164-10-192. BMC Genomics. 2009. PMID: 19393068 Free PMC article.
-
Genomic characteristics of the T cell receptor (TRB) locus in the rabbit (Oryctolagus cuniculus) revealed by comparative and phylogenetic analyses.Immunogenetics. 2014 Apr;66(4):255-66. doi: 10.1007/s00251-013-0754-1. Epub 2014 Feb 6. Immunogenetics. 2014. PMID: 24500788
-
Characterization of the ferret TRB locus guided by V, D, J, and C gene expression analysis.Immunogenetics. 2020 Feb;72(1-2):101-108. doi: 10.1007/s00251-019-01142-9. Epub 2019 Dec 3. Immunogenetics. 2020. PMID: 31797007 Free PMC article. Review.
-
The Camel Adaptive Immune Receptors Repertoire as a Singular Example of Structural and Functional Genomics.Front Genet. 2019 Oct 17;10:997. doi: 10.3389/fgene.2019.00997. eCollection 2019. Front Genet. 2019. PMID: 31681428 Free PMC article. Review.
Cited by
-
Different Species of Bats: Genomics, Transcriptome, and Immune Repertoire.Curr Issues Mol Biol. 2025 Apr 7;47(4):252. doi: 10.3390/cimb47040252. Curr Issues Mol Biol. 2025. PMID: 40699651 Free PMC article. Review.
-
Evolution of T cell receptor beta loci in salmonids.Front Immunol. 2023 Aug 15;14:1238321. doi: 10.3389/fimmu.2023.1238321. eCollection 2023. Front Immunol. 2023. PMID: 37649482 Free PMC article.
-
IMGT® Biocuration and Comparative Study of the T Cell Receptor Beta Locus of Veterinary Species Based on Homo sapiens TRB.Front Immunol. 2020 May 5;11:821. doi: 10.3389/fimmu.2020.00821. eCollection 2020. Front Immunol. 2020. PMID: 32431713 Free PMC article.
-
The Genomic Organisation of the TRA/TRD Locus Validates the Peculiar Characteristics of Dromedary δ-Chain Expression.Genes (Basel). 2021 Apr 9;12(4):544. doi: 10.3390/genes12040544. Genes (Basel). 2021. PMID: 33918850 Free PMC article.
-
Evolution of the T-Cell Receptor (TR) Loci in the Adaptive Immune Response: The Tale of the TRG Locus in Mammals.Genes (Basel). 2020 Jun 5;11(6):624. doi: 10.3390/genes11060624. Genes (Basel). 2020. PMID: 32517024 Free PMC article. Review.
References
-
- Lefranc M, Lefranc GP. The T cell Receptor Facts-Book. (New York, NY: Academic; ) (2001). p. 1–398.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

