Draft genome sequence and detailed characterization of biofuel production by oleaginous microalga Scenedesmus quadricauda LWG002611
- PMID: 30455737
- PMCID: PMC6225629
- DOI: 10.1186/s13068-018-1308-4
Draft genome sequence and detailed characterization of biofuel production by oleaginous microalga Scenedesmus quadricauda LWG002611
Abstract
Background: Due to scarcity of fossil fuel, the importance of alternative energy sources is ever increasing. The oleaginous microalgae have demonstrated their potential as an alternative source of energy, but have not achieved commercialization owing to some biological and technical inefficiency. Modern methods of recombinant strain development for improved efficacy are suffering due to inadequate knowledge of genome and limited molecular tools available for their manipulation.
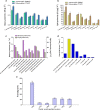
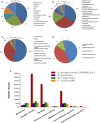
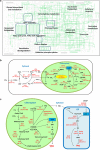
Results: In the present study, microalga Scenedesmus quadricauda LWG002611 was selected as the preferred organism for lipid production as it contained high biomass (0.37 g L-1 day-1) and lipid (102 mg L-1 day-1), compared to other oleaginous algae examined in the present study as well as earlier reports. It possessed suitable biodiesel properties as per the range defined by the European biodiesel standard EN14214 and petro-diesel standard EN590:2013. To investigate the potential of S. quadricauda LWG002611 in details, the genome of the organism was assembled and annotated. This was the first genome sequencing and assembly of S. quadricauda, which predicted a genome size of 65.35 Mb with 13,514 genes identified by de novo and 16,739 genes identified by reference guided annotation. Comparative genomics revealed that it belongs to class Chlorophyceae and order Sphaeropleales. Further, small subunit ribosomal RNA gene (18S rRNA) sequencing was carried out to confirm its molecular identification. S. quadricauda LWG002611 exhibited higher number of genes related to major activities compared to other potential algae reported earlier with a total of 283 genes identified in lipid metabolism. Metabolic pathways were reconstructed and multiple gene homologs responsible for carbon fixation and triacylglycerol (TAG) biosynthesis pathway were identified to further improve this potential algal strain for biofuel production by metabolic engineering approaches.
Conclusion: Here we present the first draft genome sequence, genetic characterization and comparative evaluation of S. quadricauda LWG002611 which exhibit high biomass as well as high lipid productivity. The knowledge of genome sequence, reconstructed metabolic pathways and identification of rate-limiting steps in TAG biosynthesis pathway will strengthen the development of molecular tools towards further improving this potentially one of the major algal strains for biofuel production.
Keywords: Biofuel; Draft genome sequence; Lipid metabolism; Metabolic pathways; Oleaginous microalgae; Phylogenetic analysis; Scenedesmus quadricauda.
Figures

Similar articles
-
The growth and lipid accumulation of Scenedesmus quadricauda during batch mixotrophic/heterotrophic cultivation using xylose as a carbon source.Bioresour Technol. 2018 Sep;263:525-531. doi: 10.1016/j.biortech.2018.05.020. Epub 2018 May 10. Bioresour Technol. 2018. PMID: 29778023
-
Genome sequencing, assembly, and annotation of the self-flocculating microalga Scenedesmus obliquus AS-6-11.BMC Genomics. 2020 Oct 27;21(1):743. doi: 10.1186/s12864-020-07142-4. BMC Genomics. 2020. PMID: 33109102 Free PMC article.
-
De novo transcriptomic analysis of an oleaginous microalga: pathway description and gene discovery for production of next-generation biofuels.PLoS One. 2012;7(4):e35142. doi: 10.1371/journal.pone.0035142. Epub 2012 Apr 20. PLoS One. 2012. Retraction in: PLoS One. 2012;7(6). doi: 10.1371/annotation/3155a3e9-5fbe-435c-a07a-e9a4846ec0b6. PMID: 22536352 Free PMC article. Retracted.
-
Microalgal lipids biochemistry and biotechnological perspectives.Biotechnol Adv. 2014 Dec;32(8):1476-93. doi: 10.1016/j.biotechadv.2014.10.003. Epub 2014 Oct 14. Biotechnol Adv. 2014. PMID: 25449285 Review.
-
A comprehensive review on carbon source effect of microalgae lipid accumulation for biofuel production.Sci Total Environ. 2022 Feb 1;806(Pt 3):151387. doi: 10.1016/j.scitotenv.2021.151387. Epub 2021 Nov 2. Sci Total Environ. 2022. PMID: 34740661 Review.
Cited by
-
Genome analyses provide insights into the evolution and adaptation of the eukaryotic Picophytoplankton Mychonastes homosphaera.BMC Genomics. 2020 Jul 11;21(1):477. doi: 10.1186/s12864-020-06891-6. BMC Genomics. 2020. PMID: 32652928 Free PMC article.
-
Environmental transcriptomics under heat stress: Can environmental RNA reveal changes in gene expression of aquatic organisms?Mol Ecol. 2025 Jul;34(13):e17152. doi: 10.1111/mec.17152. Epub 2023 Oct 4. Mol Ecol. 2025. PMID: 37792902 Free PMC article.
-
A multi-omic characterization of temperature stress in a halotolerant Scenedesmus strain for algal biotechnology.Commun Biol. 2021 Mar 12;4(1):333. doi: 10.1038/s42003-021-01859-y. Commun Biol. 2021. PMID: 33712730 Free PMC article.
-
Role of nanobionics to improve the photosynthetic productivity in plants and algae: an emerging approach.3 Biotech. 2025 Apr;15(4):74. doi: 10.1007/s13205-025-04244-2. Epub 2025 Mar 6. 3 Biotech. 2025. PMID: 40060293 Review.
-
Grand Challenges in Microalgae Domestication.Front Plant Sci. 2021 Sep 23;12:764573. doi: 10.3389/fpls.2021.764573. eCollection 2021. Front Plant Sci. 2021. PMID: 34630500 Free PMC article. No abstract available.
References
-
- Demirbas MF. Biofuels from algae for sustainable development. Appl Energy. 2011;88:3473–3480. doi: 10.1016/j.apenergy.2011.01.059. - DOI
-
- Rawat I, Kumar RR, Mutanda T, Bux F. Biodiesel from microalgae: a critical evaluation from laboratory to large scale production. Appl Energy. 2013;103:444–467. doi: 10.1016/j.apenergy.2012.10.004. - DOI
-
- Ren HY, Liu BF, Ma C, Zhao L, Ren NQ. A new lipid-rich microalga Scenedesmus sp. strain R-16 isolated using Nile red staining: effects of carbon and nitrogen sources and initial pH on the biomass and lipid production. Biotechnol Biofuels. 2013;6(1):143. doi: 10.1186/1754-6834-6-143. - DOI - PMC - PubMed
-
- Mata MT, Melo AC, Meireles S, Mendes AM, Martins AA, Caetano NS. Potential of microalgae Scenedesmus obliquus grown in brewery wastewater for biodiesel production. Chem Eng Trans. 2013;32:901–905.
LinkOut - more resources
Full Text Sources
Miscellaneous

