Using Bayes model averaging to leverage both gene main effects and G × E interactions to identify genomic regions in genome-wide association studies
- PMID: 30456811
- PMCID: PMC6375769
- DOI: 10.1002/gepi.22171
Using Bayes model averaging to leverage both gene main effects and G × E interactions to identify genomic regions in genome-wide association studies
Abstract
Genome-wide association studies typically search for marginal associations between a single-nucleotide polymorphism (SNP) and a disease trait while gene-environment (G × E) interactions remain generally unexplored. More powerful methods beyond the simple case-control (CC) approach leverage either marginal effects or CC ascertainment to increase power. However, these potential gains depend on assumptions whose aptness is often unclear a priori. Here, we review G × E methods and use simulations to highlight performance as a function of main and interaction effects and the association of the two factors in the source population. Substantial variation in performance between methods leads to uncertainty as to which approach is most appropriate for any given analysis. We present a framework that (a) balances the robustness of a CC approach with the power of the case-only (CO) approach; (b) incorporates main SNP effects; (c) allows for incorporation of prior information; and (d) allows the data to determine the most appropriate model. Our framework is based on Bayes model averaging, which provides a principled statistical method for incorporating model uncertainty. We average over inclusion of parameters corresponding to the main and G × E interaction effects and the G-E association in controls. The resulting method exploits the joint evidence for main and interaction effects while gaining power from a CO equivalent analysis. Through simulations, we demonstrate that our approach detects SNPs within a wide range of scenarios with increased power over current methods. We illustrate the approach on a gene-environment scan in the USC Children's Health Study.
Keywords: bayesian model; case-control studies; environmental factor; genome-wide scan; power.
© 2018 Wiley Periodicals, Inc.
Figures
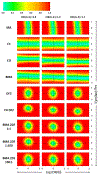



References
-
- Agresti A (2002). Loglinear Models for Contingency Tables Categorical Data Analysis (2 ed., pp. 314–356). New Jersey: John Wiley & Sons.
-
- Bishop YMM, Fienberg SE, & Holland PW (1975). Discrete multivariate analysis : theory and practice. Cambridge, Mass. ; London: M.I.T. Press.
-
- Hoeting JA, Madigan D, Raftery AE, & Volinsky CT (1999). Bayesian model averaging: A tutorial. Statistical Science, 14(4), 382–401.
Publication types
MeSH terms
Substances
Grants and funding
- R01CA201407/NH/NIH HHS/United States
- P30 ES007048/ES/NIEHS NIH HHS/United States
- R01CA140561/NH/NIH HHS/United States
- R01ES016813/NH/NIH HHS/United States
- T32ES013678/ES/NIEHS NIH HHS/United States
- P01CA196569/NH/NIH HHS/United States
- U01 CA188392/CA/NCI NIH HHS/United States
- R01 ES016813/ES/NIEHS NIH HHS/United States
- R01 CA140561/CA/NCI NIH HHS/United States
- P30ES007048/ES/NIEHS NIH HHS/United States
- R01 CA201407/CA/NCI NIH HHS/United States
- T32 ES013678/ES/NIEHS NIH HHS/United States
- R21 ES024844/ES/NIEHS NIH HHS/United States
- P01 CA196569/CA/NCI NIH HHS/United States
- ES024844/NH/NIH HHS/United States
LinkOut - more resources
Full Text Sources

