Anti-EGFR-resistant clones decay exponentially after progression: implications for anti-EGFR re-challenge
- PMID: 30462160
- PMCID: PMC6657008
- DOI: 10.1093/annonc/mdy509
Anti-EGFR-resistant clones decay exponentially after progression: implications for anti-EGFR re-challenge
Abstract
Background: Colorectal cancer (CRC) has been shown to acquire RAS and EGFR ectodomain mutations as mechanisms of resistance to epidermal growth factor receptor (EGFR) inhibition (anti-EGFR). After anti-EGFR withdrawal, RAS and EGFR mutant clones lack a growth advantage relative to other clones and decay; however, the kinetics of decay remain unclear. We sought to determine the kinetics of acquired RAS/EGFR mutations after discontinuation of anti-EGFR therapy.
Patients and methods: We present the post-progression circulating tumor DNA (ctDNA) profiles of 135 patients with RAS/BRAF wild-type metastatic CRC treated with anti-EGFR who acquired RAS and/or EGFR mutations during therapy. Our validation cohort consisted of an external dataset of 73 patients with a ctDNA profile suggestive of prior anti-EGFR exposure and serial sampling. A separate retrospective cohort of 80 patients was used to evaluate overall response rate and progression free survival during re-challenge therapies.
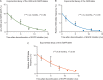
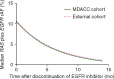
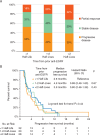
Results: Our analysis showed that RAS and EGFR relative mutant allele frequency decays exponentially (r2=0.93 for RAS; r2=0.94 for EGFR) with a cumulative half-life of 4.4 months. We validated our findings using an external dataset of 73 patients with a ctDNA profile suggestive of prior anti-EGFR exposure and serial sampling, confirming exponential decay with an estimated half-life of 4.3 months. A separate retrospective cohort of 80 patients showed that patients had a higher overall response rate during re-challenge therapies after increasing time intervals, as predicted by our model.
Conclusion: These results provide scientific support for anti-EGFR re-challenge and guide the optimal timing of re-challenge initiation.
Keywords: anti-EGFR therapy; circulating tumor DNA; clonal decay; colorectal cancer.
© The Author(s) 2018. Published by Oxford University Press on behalf of the European Society for Medical Oncology. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures

Comment in
-
Failure is not final: ctDNA-guided rechallenge therapy in colorectal cancer.Ann Oncol. 2019 Feb 1;30(2):157-159. doi: 10.1093/annonc/mdy525. Ann Oncol. 2019. PMID: 30475974 No abstract available.
References
-
- Amado RG, Wolf M, Peeters M. et al. Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J Clin Oncol 2008; 26(10): 1626–1634. - PubMed
-
- Karapetis CS, Khambata-Ford S, Jonker DJ. et al. K-ras mutations and benefit from cetuximab in advanced colorectal cancer. N Engl J Med 2008; 359(17): 1757–1765. - PubMed
-
- Douillard JY, Oliner KS, Siena S. et al. Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer. N Engl J Med 2013; 369(11): 1023–1034. - PubMed
-
- Van Cutsem E, Kohne CH, Hitre E. et al. Cetuximab and chemotherapy as initial treatment for metastatic colorectal cancer. N Engl J Med 2009; 360(14): 1408–1417. - PubMed
-
- Di Nicolantonio F, Martini M, Molinari F. et al. Wild-type BRAF is required for response to panitumumab or cetuximab in metastatic colorectal cancer. J Clin Oncol 2008; 26(35): 5705–5712. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

