The NF-κB Factor Relish Regulates Atg1 Expression and Controls Autophagy
- PMID: 30463009
- PMCID: PMC6329390
- DOI: 10.1016/j.celrep.2018.10.076
The NF-κB Factor Relish Regulates Atg1 Expression and Controls Autophagy
Abstract
Macroautophagy and cell death both contribute to innate immunity, but little is known about how these processes integrate. Drosophila larval salivary glands require autophagy for developmentally programmed cell death, and innate immune signaling factors increase in these dying cells. Here, we show that the nuclear factor κB (NF-κB) factor Relish, a component of the immune deficiency (Imd) pathway, is required for salivary gland degradation. Surprisingly, of the classic Imd pathway components, only Relish and the PGRP receptors were involved in salivary gland degradation. Significantly, Relish controls salivary gland degradation by regulating autophagy but not caspases. In addition, expression of either Relish or PGRP-LC causes premature autophagy induction and subsequent gland degradation. Relish controls autophagy by regulating the expression of Atg1, a core component and activator of the autophagy pathway. Together these findings demonstrate that a NF-κB pathway regulates autophagy during developmentally programmed cell death.
Keywords: Drosophila; NF-κB; autophagy; cell death.
Copyright © 2018 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
DECLARATION OF INTERESTS
The authors declare no competing interests.
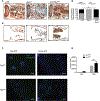
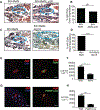
Figures




Similar articles
-
Activation of autophagy during cell death requires the engulfment receptor Draper.Nature. 2010 Jun 24;465(7301):1093-6. doi: 10.1038/nature09127. Nature. 2010. PMID: 20577216 Free PMC article.
-
The engulfment receptor Draper is required for autophagy during cell death.Autophagy. 2010 Nov;6(8):1192-3. doi: 10.4161/auto.6.8.13474. Epub 2010 Nov 16. Autophagy. 2010. PMID: 20864812 Free PMC article.
-
Drosophila H2Av negatively regulates the activity of the IMD pathway via facilitating Relish SUMOylation.PLoS Genet. 2021 Aug 9;17(8):e1009718. doi: 10.1371/journal.pgen.1009718. eCollection 2021 Aug. PLoS Genet. 2021. PMID: 34370736 Free PMC article.
-
Dynamic Regulation of NF-κB Response in Innate Immunity: The Case of the IMD Pathway in Drosophila.Biomedicines. 2022 Sep 16;10(9):2304. doi: 10.3390/biomedicines10092304. Biomedicines. 2022. PMID: 36140409 Free PMC article. Review.
-
Autophagy in Drosophila melanogaster.Biochim Biophys Acta. 2009 Sep;1793(9):1452-60. doi: 10.1016/j.bbamcr.2009.02.009. Epub 2009 Mar 2. Biochim Biophys Acta. 2009. PMID: 19264097 Free PMC article. Review.
Cited by
-
Yorkie Growth-Promoting Activity Is Limited by Atg1-Mediated Phosphorylation.Dev Cell. 2020 Mar 9;52(5):605-616.e7. doi: 10.1016/j.devcel.2020.01.011. Epub 2020 Feb 6. Dev Cell. 2020. PMID: 32032548 Free PMC article.
-
Autophagic dysfunction and gut microbiota dysbiosis cause chronic immune activation in a Drosophila model of Gaucher disease.PLoS Genet. 2023 Dec 21;19(12):e1011063. doi: 10.1371/journal.pgen.1011063. eCollection 2023 Dec. PLoS Genet. 2023. PMID: 38127816 Free PMC article.
-
Transcription and Post-translational Regulation of Autophagy in Insects.Front Physiol. 2022 Feb 25;13:825202. doi: 10.3389/fphys.2022.825202. eCollection 2022. Front Physiol. 2022. PMID: 35283796 Free PMC article.
-
Regulation of the Drosophila Imd pathway by signaling amyloids.Insect Biochem Mol Biol. 2019 May;108:16-23. doi: 10.1016/j.ibmb.2019.03.003. Epub 2019 Mar 9. Insect Biochem Mol Biol. 2019. PMID: 30857831 Free PMC article. Review.
-
Autophagic-Related Proteins in Brain Gliomas: Role, Mechanisms, and Targeting Agents.Cancers (Basel). 2023 May 5;15(9):2622. doi: 10.3390/cancers15092622. Cancers (Basel). 2023. PMID: 37174088 Free PMC article. Review.
References
-
- Andres AJ, and Thummel CS (1994). Methods for quantitative analysis of transcription in larvae and prepupae. Methods Cell Biol 44, 565–573. - PubMed
-
- Bonizzi G, and Karin M (2004). The two NF-kappaB activation pathways and their role in innate and adaptive immunity. Trends Immunol 25, 280–288. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

