Molecular insights into genome-wide association studies of chronic kidney disease-defining traits
- PMID: 30467309
- PMCID: PMC6250666
- DOI: 10.1038/s41467-018-07260-4
Molecular insights into genome-wide association studies of chronic kidney disease-defining traits
Abstract
Genome-wide association studies (GWAS) have identified >100 loci of chronic kidney disease-defining traits (CKD-dt). Molecular mechanisms underlying these associations remain elusive. Using 280 kidney transcriptomes and 9958 gene expression profiles from 44 non-renal tissues we uncover gene expression partners (eGenes) for 88.9% of CKD-dt GWAS loci. Through epigenomic chromatin segmentation analysis and variant effect prediction we annotate functional consequences to 74% of these loci. Our colocalisation analysis and Mendelian randomisation in >130,000 subjects demonstrate causal effects of three eGenes (NAT8B, CASP9 and MUC1) on estimated glomerular filtration rate. We identify a common alternative splice variant in MUC1 (a gene responsible for rare Mendelian form of kidney disease) and observe increased renal expression of a specific MUC1 mRNA isoform as a plausible molecular mechanism of the GWAS association signal. These data highlight the variants and genes underpinning the associations uncovered in GWAS of CKD-dt.
Conflict of interest statement
The authors declare no competing interests.
Figures
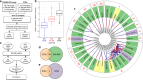


References
-
- Levin, A. et al. Global kidney health 2017 and beyond: a roadmap for closing gaps in care, research, and policy. Lancet390, 1888–1917 (2017). - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

