Automated and flexible identification of complex disease: building a model for systemic lupus erythematosus using noisy labeling
- PMID: 30476175
- PMCID: PMC6308013
- DOI: 10.1093/jamia/ocy154
Automated and flexible identification of complex disease: building a model for systemic lupus erythematosus using noisy labeling
Abstract
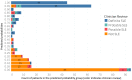
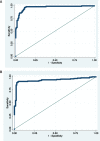
Accurate and efficient identification of complex chronic conditions in the electronic health record (EHR) is an important but challenging task that has historically relied on tedious clinician review and oversimplification of the disease. Here we adapt methods that allow for automated "noisy labeling" of positive and negative controls to create a "silver standard" for machine learning to automate identification of systemic lupus erythematosus (SLE). Our final model, which includes both structured data as well as text processing of clinical notes, outperformed all existing algorithms for SLE (AUC 0.97). In addition, we demonstrate how the probabilistic outputs of this model can be adapted to various clinical needs, selecting high thresholds when specificity is the priority and lower thresholds when a more inclusive patient population is desired. Deploying a similar methodology to other complex diseases has the potential to dramatically simplify the landscape of population identification in the EHR.
Mesh terms: Electronic Health Records, Machine Learning, Lupus Erythematosus, Phenotype, Algorithms.
Figures

References
-
- Moores KG, Sathe NA.. A systematic review of validated methods for identifying systemic lupus erythematosus (SLE) using administrative or claims data. Vaccine 2013; 31: K62–73. - PubMed

