Aneuploid acute myeloid leukemia exhibits a signature of genomic alterations in the cell cycle and protein degradation machinery
- PMID: 30480765
- PMCID: PMC6587451
- DOI: 10.1002/cncr.31837
Aneuploid acute myeloid leukemia exhibits a signature of genomic alterations in the cell cycle and protein degradation machinery
Erratum in
-
Erratum.Cancer. 2021 Jun 15;127(12):2160. doi: 10.1002/cncr.33177. Epub 2021 Jan 26. Cancer. 2021. PMID: 34029393 Free PMC article. No abstract available.
Abstract
Background: Aneuploidy occurs in more than 20% of acute myeloid leukemia (AML) cases and correlates with an adverse prognosis.
Methods: To understand the molecular bases of aneuploid acute myeloid leukemia (A-AML), this study examined the genomic profile in 42 A-AML cases and 35 euploid acute myeloid leukemia (E-AML) cases.
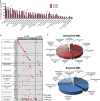
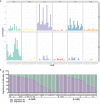
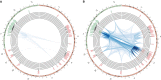
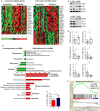
Results: A-AML was characterized by increased genomic complexity based on exonic variants (an average of 26 somatic mutations per sample vs 15 for E-AML). The integration of exome, copy number, and gene expression data revealed alterations in genes involved in DNA repair (eg, SLX4IP, RINT1, HINT1, and ATR) and the cell cycle (eg, MCM2, MCM4, MCM5, MCM7, MCM8, MCM10, UBE2C, USP37, CK2, CK3, CK4, BUB1B, NUSAP1, and E2F) in A-AML, which was associated with a 3-gene signature defined by PLK1 and CDC20 upregulation and RAD50 downregulation and with structural or functional silencing of the p53 transcriptional program. Moreover, A-AML was enriched for alterations in the protein ubiquitination and degradation pathway (eg, increased levels of UHRF1 and UBE2C and decreased UBA3 expression), response to reactive oxygen species, energy metabolism, and biosynthetic processes, which may help in facing the unbalanced protein load. E-AML was associated with BCOR/BCORL1 mutations and HOX gene overexpression.
Conclusions: These findings indicate that aneuploidy-related and leukemia-specific alterations cooperate to tolerate an abnormal chromosome number in AML, and they point to the mitotic and protein degradation machineries as potential therapeutic targets.
Keywords: acute myeloid leukemia; aneuploidy; cell cycle; genomics; mutation; ubiquitination; whole exome sequencing.
© 2018 The Authors. Cancer published by Wiley Periodicals, Inc. on behalf of American Cancer Society.
Conflict of interest statement
Lars Bullinger reports personal fees and nonfinancial support from Bristol‐Myers Squibb; personal fees from Novartis, Jazz Pharmaceuticals, and Pfizer; grants and personal fees from Sanofi; and nonfinancial support from Amgen outside the submitted work. Michele Cavo reports personal fees from Janssen, Celgene, Bristol‐Myers Squibb, Amgen, and Takeda and other from Novartis outside the submitted work. Torsten Haferlach reports partial ownership of the Munich Leukemia Laboratory. Giovanni Martinelli reports compensation or nonfinancial support from Amgen (consulting or advisory role), Ariad/Incyte (consulting or advisory role), Pfizer (consulting or advisory role and speakers’ bureau), Celgene (consulting or advisory role and speakers’ bureau), Janssen (consulting or advisory role), Jazz Pharmaceuticals (consulting or advisory role), AbbVie (consulting or advisory role), Novartis (speakers’ bureau), Daiichi Sankyo (travel), Shire (travel), J&J, and Roche (consulting or advisory role and travel) outside the submitted work.
Figures

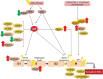
Similar articles
-
Whole-exome sequencing identifies somatic mutations of BCOR in acute myeloid leukemia with normal karyotype.Blood. 2011 Dec 1;118(23):6153-63. doi: 10.1182/blood-2011-07-365320. Epub 2011 Oct 19. Blood. 2011. PMID: 22012066
-
Integrated genomics of susceptibility to alkylator-induced leukemia in mice.BMC Genomics. 2010 Nov 17;11:638. doi: 10.1186/1471-2164-11-638. BMC Genomics. 2010. PMID: 21080971 Free PMC article.
-
Identification of somatic mutations using whole-exome sequencing in Korean patients with acute myeloid leukemia.BMC Med Genet. 2017 Mar 1;18(1):23. doi: 10.1186/s12881-017-0382-y. BMC Med Genet. 2017. PMID: 28249600 Free PMC article.
-
What do functional genomics tell us about pathogenesis of AML?Best Pract Res Clin Haematol. 2019 Dec;32(4):101101. doi: 10.1016/j.beha.2019.101101. Epub 2019 Oct 18. Best Pract Res Clin Haematol. 2019. PMID: 31779979 Review.
-
[Identification of novel pathogenic gene mutations in pediatric acute myeloid leukemia by whole-exome resequencing].Rinsho Ketsueki. 2015 Dec;56(12):2419-25. doi: 10.11406/rinketsu.56.2419. Rinsho Ketsueki. 2015. PMID: 26725349 Review. Japanese.
Cited by
-
Targeting Proliferation Signals and the Cell Cycle Machinery in Acute Leukemias: Novel Molecules on the Horizon.Molecules. 2023 Jan 26;28(3):1224. doi: 10.3390/molecules28031224. Molecules. 2023. PMID: 36770891 Free PMC article. Review.
-
MRE11-RAD50-NBS1 complex alterations and DNA damage response: implications for cancer treatment.Mol Cancer. 2019 Nov 26;18(1):169. doi: 10.1186/s12943-019-1100-5. Mol Cancer. 2019. PMID: 31767017 Free PMC article. Review.
-
RINT1 Loss Impairs Retinogenesis Through TRP53-Mediated Apoptosis.Front Cell Dev Biol. 2020 Jul 30;8:711. doi: 10.3389/fcell.2020.00711. eCollection 2020. Front Cell Dev Biol. 2020. PMID: 32850831 Free PMC article.
-
lncRNA lnc-POP1-1 upregulated by VN1R5 promotes cisplatin resistance in head and neck squamous cell carcinoma through interaction with MCM5.Mol Ther. 2022 Jan 5;30(1):448-467. doi: 10.1016/j.ymthe.2021.06.006. Epub 2021 Jun 8. Mol Ther. 2022. PMID: 34111560 Free PMC article.
-
Feasibility of Leukemia-Derived Exosome Enrichment and Co-isolated dsDNA Sequencing in Acute Myeloid Leukemia Patients: A Proof of Concept for New Leukemia Biomarkers Detection.Cancers (Basel). 2022 Sep 16;14(18):4504. doi: 10.3390/cancers14184504. Cancers (Basel). 2022. PMID: 36139664 Free PMC article.
References
-
- Torres EM, Sokolsky T, Tucker CM, et al. Effects of aneuploidy on cellular physiology and cell division in haploid yeast. Science. 2007;317:916‐924. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

