Role of the uS9/yS16 C-terminal tail in translation initiation and elongation in Saccharomyces cerevisiae
- PMID: 30481328
- PMCID: PMC6344880
- DOI: 10.1093/nar/gky1180
Role of the uS9/yS16 C-terminal tail in translation initiation and elongation in Saccharomyces cerevisiae
Abstract
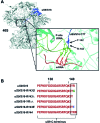
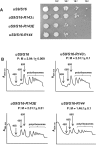
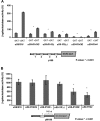
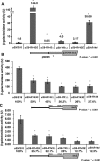
The small ribosomal subunit protein uS9 (formerly called rpS16 in Saccharomyces cerevisiae), has a long protruding C-terminal tail (CTT) that extends towards the mRNA cleft of the ribosome. The last C-terminal residue of uS9 is an invariably conserved, positively charged Arg that is believed to enhance interaction of the negatively charged initiator tRNA with the ribosome when the tRNA is base-paired to the AUG codon in the P-site. In order to more fully characterize the role of the uS9 CTT in eukaryotic translation, we tested how truncations, extensions and substitutions within the CTT affect initiation and elongation processes in Saccharomyces cerevisiae. We found that uS9 C-terminal residues are critical for efficient recruitment of the eIF2•GTP•Met-tRNAiMet ternary complex to the ribosome and for its proper response to the presence of an AUG codon in the P-site during the scanning phase of initiation. These residues also regulate hydrolysis of the GTP in the eIF2•GTP•Met-tRNAiMet complex to GDP and Pi. In addition, our data show that uS9 CTT modulates elongation fidelity. Therefore, we propose that uS9 CTT is critical for proper control of the complex interplay of events surrounding accommodation of initiator and elongator tRNAs in the P- and A-sites of the ribosome.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

