The Different Tactics of Foot-and-Mouth Disease Virus to Evade Innate Immunity
- PMID: 30483224
- PMCID: PMC6241212
- DOI: 10.3389/fmicb.2018.02644
The Different Tactics of Foot-and-Mouth Disease Virus to Evade Innate Immunity
Abstract
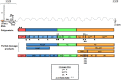
Like all pathogens, foot-and-mouth disease virus (FMDV) is recognized by the immune system inducing a heightened immune response mainly mediated by type I and type III IFNs. To overcome the strong antiviral response induced by these cytokines, FMDV has evolved many strategies exploiting each region of its small RNA genome. These include: (a) inhibition of IFN induction at the transcriptional and translational level, (b) inhibition of protein trafficking; (c) blockage of specific post-translational modifications in proteins that regulate innate immune signaling; (d) modulation of autophagy; (e) inhibition of stress granule formation; and (f) in vivo modulation of immune cell function. Here, we summarize and discuss FMDV virulence factors and the host immune footprint that characterize infection in cell culture and in the natural hosts.
Keywords: FMDV; IFN; apoptosis; autophagy; cattle/swine; evasion innate immunity; ubiquitin/ISG; virulence factors.
Figures

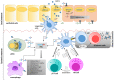
References
Publication types
LinkOut - more resources
Full Text Sources

